+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8ivq | ||||||
|---|---|---|---|---|---|---|---|
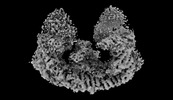
| Title | Cryo-EM structure of mouse BIRC6, Global map | ||||||
 Components Components | Isoform 2 of Baculoviral IAP repeat-containing protein 6 | ||||||
 Keywords Keywords |  APOPTOSIS / APOPTOSIS /  Inhibitor of apoptosis protein / BIRC6 / Smac Inhibitor of apoptosis protein / BIRC6 / Smac | ||||||
| Function / homology |  Function and homology information Function and homology informationspongiotrophoblast layer development / labyrinthine layer development / Flemming body /  microtubule organizing center / cysteine-type endopeptidase inhibitor activity / ubiquitin conjugating enzyme activity / placenta development / microtubule organizing center / cysteine-type endopeptidase inhibitor activity / ubiquitin conjugating enzyme activity / placenta development /  regulation of cytokinesis / negative regulation of extrinsic apoptotic signaling pathway / RING-type E3 ubiquitin transferase ...spongiotrophoblast layer development / labyrinthine layer development / Flemming body / regulation of cytokinesis / negative regulation of extrinsic apoptotic signaling pathway / RING-type E3 ubiquitin transferase ...spongiotrophoblast layer development / labyrinthine layer development / Flemming body /  microtubule organizing center / cysteine-type endopeptidase inhibitor activity / ubiquitin conjugating enzyme activity / placenta development / microtubule organizing center / cysteine-type endopeptidase inhibitor activity / ubiquitin conjugating enzyme activity / placenta development /  regulation of cytokinesis / negative regulation of extrinsic apoptotic signaling pathway / RING-type E3 ubiquitin transferase / regulation of cytokinesis / negative regulation of extrinsic apoptotic signaling pathway / RING-type E3 ubiquitin transferase /  trans-Golgi network / trans-Golgi network /  spindle pole / ubiquitin-protein transferase activity / regulation of cell population proliferation / midbody / cell population proliferation / protein ubiquitination / spindle pole / ubiquitin-protein transferase activity / regulation of cell population proliferation / midbody / cell population proliferation / protein ubiquitination /  endosome / endosome /  cell cycle / cell cycle /  cell division / apoptotic process / positive regulation of cell population proliferation / negative regulation of apoptotic process / cell division / apoptotic process / positive regulation of cell population proliferation / negative regulation of apoptotic process /  membrane / membrane /  nucleus nucleusSimilarity search - Function | ||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | ||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.6 Å cryo EM / Resolution: 3.6 Å | ||||||
 Authors Authors | Liu, S. / Jiang, T. / Bu, F. / Zhao, J. / Wang, G. / Li, N. / Gao, N. / Qiu, X. | ||||||
| Funding support |  China, 1items China, 1items
| ||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Molecular mechanisms underlying the BIRC6-mediated regulation of apoptosis and autophagy. Authors: Shuo-Shuo Liu / Tian-Xia Jiang / Fan Bu / Ji-Lan Zhao / Guang-Fei Wang / Guo-Heng Yang / Jie-Yan Kong / Yun-Fan Qie / Pei Wen / Li-Bin Fan / Ning-Ning Li / Ning Gao / Xiao-Bo Qiu /  Abstract: Procaspase 9 is the initiator caspase for apoptosis, but how its levels and activities are maintained remains unclear. The gigantic Inhibitor-of-Apoptosis Protein BIRC6/BRUCE/Apollon inhibits both ...Procaspase 9 is the initiator caspase for apoptosis, but how its levels and activities are maintained remains unclear. The gigantic Inhibitor-of-Apoptosis Protein BIRC6/BRUCE/Apollon inhibits both apoptosis and autophagy by promoting ubiquitylation of proapoptotic factors and the key autophagic protein LC3, respectively. Here we show that BIRC6 forms an anti-parallel U-shaped dimer with multiple previously unannotated domains, including a ubiquitin-like domain, and the proapoptotic factor Smac/DIABLO binds BIRC6 in the central cavity. Notably, Smac outcompetes the effector caspase 3 and the pro-apoptotic protease HtrA2, but not procaspase 9, for binding BIRC6 in cells. BIRC6 also binds LC3 through its LC3-interacting region, probably following dimer disruption of this BIRC6 region. Mutation at LC3 ubiquitylation site promotes autophagy and autophagic degradation of BIRC6. Moreover, induction of autophagy promotes autophagic degradation of BIRC6 and caspase 9, but not of other effector caspases. These results are important to understand how the balance between apoptosis and autophagy is regulated under pathophysiological conditions. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8ivq.cif.gz 8ivq.cif.gz | 1 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8ivq.ent.gz pdb8ivq.ent.gz | Display |  PDB format PDB format | |
| PDBx/mmJSON format |  8ivq.json.gz 8ivq.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/iv/8ivq https://data.pdbj.org/pub/pdb/validation_reports/iv/8ivq ftp://data.pdbj.org/pub/pdb/validation_reports/iv/8ivq ftp://data.pdbj.org/pub/pdb/validation_reports/iv/8ivq | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  35759MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 535020.812 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Details: The plasmid used for BIRC6 protein expression in this deposition is originated from Dr. Stefan Jentsch lab, and the Birc6 sequence in that plasmid is available from GenBank under accession ...Details: The plasmid used for BIRC6 protein expression in this deposition is originated from Dr. Stefan Jentsch lab, and the Birc6 sequence in that plasmid is available from GenBank under accession number GenBank: Y17267.1. No identical sequence could be identified in Uniprot. Here is the citation of the used Birc6 sequence: Hauser, H.P., et al., A giant ubiquitin-conjugating enzyme related to IAP apoptosis inhibitors. Journal of Cell Biology, 1998. 141(6): p. 1415-1422. Source: (gene. exp.)   Mus musculus (house mouse) / Gene: Birc6, Kiaa1289 / Production host: Mus musculus (house mouse) / Gene: Birc6, Kiaa1289 / Production host:   Homo sapiens (human) Homo sapiens (human)References: UniProt: O88738, RING-type E3 ubiquitin transferase |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Baculoviral IAP repeat-containing protein 6, Dimer / Type: COMPLEX / Entity ID: all / Source: RECOMBINANT |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
| Source (recombinant) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Buffer solution | pH: 7.5 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 3000 nm / Nominal defocus min: 2000 nm Bright-field microscopy / Nominal defocus max: 3000 nm / Nominal defocus min: 2000 nm |
| Image recording | Electron dose: 68.4 e/Å2 / Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) |
- Processing
Processing
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
3D reconstruction | Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 154000 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller







 PDBj
PDBj


