[English] 日本語
 Yorodumi
Yorodumi- PDB-7tpj: Single-Particle Cryo-EM Structure of the WaaL O-antigen ligase in... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7tpj | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
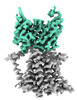
| Title | Single-Particle Cryo-EM Structure of the WaaL O-antigen ligase in its apo state | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 Components Components |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 Keywords Keywords |  MEMBRANE PROTEIN / MEMBRANE PROTEIN /  Lipopolysaccharide / single-particle cryo-electron microscopy / molecular dynamics simulations / Lipopolysaccharide / single-particle cryo-electron microscopy / molecular dynamics simulations /  structural biology / structural biology /  undecaprenyl pyrophosphate / WaaL Ligase / undecaprenyl pyrophosphate / WaaL Ligase /  lipid A / O-antigen / lipid A / O-antigen /  membrane proteins / transglycosylation / membrane proteins / transglycosylation /  glycosyltransferase glycosyltransferase | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Function / homology | O-antigen ligase-related / O-Antigen ligase /  ligase activity / membrane => GO:0016020 / Putative cell surface polysaccharide polymerase/ligase ligase activity / membrane => GO:0016020 / Putative cell surface polysaccharide polymerase/ligase Function and homology information Function and homology information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological species |   Cupriavidus metallidurans (bacteria) Cupriavidus metallidurans (bacteria)  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.46 Å cryo EM / Resolution: 3.46 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 Authors Authors | Ashraf, K.U. / Nygaard, R. / Vickery, O.N. / Erramilli, S.K. / Herrera, C.M. / McConville, T.H. / Petrou, V.I. / Giacometti, S.I. / Dufrisne, M.B. / Nosol, K. ...Ashraf, K.U. / Nygaard, R. / Vickery, O.N. / Erramilli, S.K. / Herrera, C.M. / McConville, T.H. / Petrou, V.I. / Giacometti, S.I. / Dufrisne, M.B. / Nosol, K. / Zinkle, A.P. / Graham, C.L.B. / Loukeris, M. / Kloss, B. / Skorupinska-Tudek, K. / Swiezewska, E. / Roper, D. / Clarke, O.B. / Uhlemann, A.C. / Kossiakoff, A.A. / Trent, M.S. / Stansfeld, P.J. / Mancia, F. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Funding support |  United States, United States,  United Kingdom, 18items United Kingdom, 18items
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Structural basis of lipopolysaccharide maturation by the O-antigen ligase. Authors: Khuram U Ashraf / Rie Nygaard / Owen N Vickery / Satchal K Erramilli / Carmen M Herrera / Thomas H McConville / Vasileios I Petrou / Sabrina I Giacometti / Meagan Belcher Dufrisne / Kamil ...Authors: Khuram U Ashraf / Rie Nygaard / Owen N Vickery / Satchal K Erramilli / Carmen M Herrera / Thomas H McConville / Vasileios I Petrou / Sabrina I Giacometti / Meagan Belcher Dufrisne / Kamil Nosol / Allen P Zinkle / Chris L B Graham / Michael Loukeris / Brian Kloss / Karolina Skorupinska-Tudek / Ewa Swiezewska / David I Roper / Oliver B Clarke / Anne-Catrin Uhlemann / Anthony A Kossiakoff / M Stephen Trent / Phillip J Stansfeld / Filippo Mancia /    Abstract: The outer membrane of Gram-negative bacteria has an external leaflet that is largely composed of lipopolysaccharide, which provides a selective permeation barrier, particularly against antimicrobials. ...The outer membrane of Gram-negative bacteria has an external leaflet that is largely composed of lipopolysaccharide, which provides a selective permeation barrier, particularly against antimicrobials. The final and crucial step in the biosynthesis of lipopolysaccharide is the addition of a species-dependent O-antigen to the lipid A core oligosaccharide, which is catalysed by the O-antigen ligase WaaL. Here we present structures of WaaL from Cupriavidus metallidurans, both in the apo state and in complex with its lipid carrier undecaprenyl pyrophosphate, determined by single-particle cryo-electron microscopy. The structures reveal that WaaL comprises 12 transmembrane helices and a predominantly α-helical periplasmic region, which we show contains many of the conserved residues that are required for catalysis. We observe a conserved fold within the GT-C family of glycosyltransferases and hypothesize that they have a common mechanism for shuttling the undecaprenyl-based carrier to and from the active site. The structures, combined with genetic, biochemical, bioinformatics and molecular dynamics simulation experiments, offer molecular details on how the ligands come in apposition, and allows us to propose a mechanistic model for catalysis. Together, our work provides a structural basis for lipopolysaccharide maturation in a member of the GT-C superfamily of glycosyltransferases. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7tpj.cif.gz 7tpj.cif.gz | 117.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7tpj.ent.gz pdb7tpj.ent.gz | 91.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7tpj.json.gz 7tpj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tp/7tpj https://data.pdbj.org/pub/pdb/validation_reports/tp/7tpj ftp://data.pdbj.org/pub/pdb/validation_reports/tp/7tpj ftp://data.pdbj.org/pub/pdb/validation_reports/tp/7tpj | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 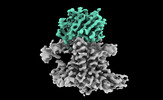 26057MC  7tpgC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 44536.754 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Cupriavidus metallidurans (bacteria) / Production host: Cupriavidus metallidurans (bacteria) / Production host:   Escherichia coli (E. coli) / References: UniProt: Q1LJU1 Escherichia coli (E. coli) / References: UniProt: Q1LJU1 |
|---|---|
| #2: Antibody | Mass: 25077.119 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| #3: Antibody | Mass: 23396.951 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Apo CmWaaL with Fab fragment bound / Type: COMPLEX / Entity ID: all / Source: MULTIPLE SOURCES | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer solution | pH: 7.5 | |||||||||||||||
| Buffer component |
| |||||||||||||||
| Specimen | Conc.: 1 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES | |||||||||||||||
Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Humidity: 95 % / Chamber temperature: 277 K |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: FEI TITAN |
|---|---|
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2000 nm / Nominal defocus min: 1000 nm Bright-field microscopy / Nominal defocus max: 2000 nm / Nominal defocus min: 1000 nm |
| Specimen holder | Cryogen: NITROGEN / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Image recording | Electron dose: 70 e/Å2 / Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Num. of grids imaged: 1 / Num. of real images: 2378 |
- Processing
Processing
| Software | Name: PHENIX / Version: 1.19_4092: / Classification: refinement | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||
CTF correction | Details: Patch CTF / Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
3D reconstruction | Resolution: 3.46 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 30514 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller


 PDBj
PDBj

