+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6ve4 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Pentadecameric PilQ from Pseudomonas aeruginosa | ||||||
 Components Components | Fimbrial assembly protein PilQ | ||||||
 Keywords Keywords |  PROTEIN TRANSPORT / PROTEIN TRANSPORT /  Type IV pilus / T4P / PilQ / Type IV pilus / T4P / PilQ /  secretin / type IVa pilus / T4aP / secretin / type IVa pilus / T4aP /  pilus / outer membrane / pilus / outer membrane /  periplasm / periplasm /  bacterial secretion system bacterial secretion system | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |   Pseudomonas aeruginosa (bacteria) Pseudomonas aeruginosa (bacteria) | ||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 6.9 Å cryo EM / Resolution: 6.9 Å | ||||||
 Authors Authors | McCallum, M. / Tammam, S. / Rubinstein, J.L. / Burrows, L.L. / Howell, P.L. | ||||||
| Funding support |  Canada, 1items Canada, 1items
| ||||||
 Citation Citation |  Journal: Structure / Year: 2021 Journal: Structure / Year: 2021Title: CryoEM map of Pseudomonas aeruginosa PilQ enables structural characterization of TsaP. Authors: Matthew McCallum / Stephanie Tammam / John L Rubinstein / Lori L Burrows / P Lynne Howell /  Abstract: The type IV pilus machinery is a multi-protein complex that polymerizes and depolymerizes a pilus fiber used for attachment, twitching motility, phage adsorption, natural competence, protein ...The type IV pilus machinery is a multi-protein complex that polymerizes and depolymerizes a pilus fiber used for attachment, twitching motility, phage adsorption, natural competence, protein secretion, and surface-sensing. An outer membrane secretin pore is required for passage of the pilus fiber out of the cell. Herein, the structure of the tetradecameric secretin, PilQ, from the Pseudomonas aeruginosa type IVa pilus system was determined to 4.3 Å and 4.4 Å resolution in the presence and absence of C symmetric spikes, respectively. The heptameric spikes were found to be two tandem C-terminal domains of TsaP. TsaP forms a belt around PilQ and while it is not essential for twitching motility, overexpression of TsaP triggers a signal cascade upstream of PilY1 leading to cyclic di-GMP up-regulation. These results resolve the identity of the spikes identified with Proteobacterial PilQ homologs and may reveal a new component of the surface-sensing cyclic di-GMP signal cascade. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6ve4.cif.gz 6ve4.cif.gz | 679.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6ve4.ent.gz pdb6ve4.ent.gz | 459.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6ve4.json.gz 6ve4.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ve/6ve4 https://data.pdbj.org/pub/pdb/validation_reports/ve/6ve4 ftp://data.pdbj.org/pub/pdb/validation_reports/ve/6ve4 ftp://data.pdbj.org/pub/pdb/validation_reports/ve/6ve4 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  21154MC  6ve2C  6ve3C C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 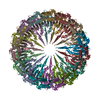
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 79731.141 Da / Num. of mol.: 15 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 Gene: pilQ, PA5040 / Production host:   Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria)Strain (production host): ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 References: UniProt: P34750 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Pentadecameric PilQ from Pseudomonas aeruginosa / Type: COMPLEX / Details: Affinity purified PilQ without TsaP / Entity ID: all / Source: RECOMBINANT |
|---|---|
| Molecular weight | Experimental value: NO |
| Source (natural) | Organism:   Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 |
| Source (recombinant) | Organism:   Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 |
| Buffer solution | pH: 8 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
| Specimen support | Details: unspecified |
Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TECNAI F20 |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Electron dose: 35.7 e/Å2 / Film or detector model: GATAN K2 SUMMIT (4k x 4k) |
- Processing
Processing
| EM software |
| |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | |||||||||||||||||||||
| Symmetry | Point symmetry : C15 (15 fold cyclic : C15 (15 fold cyclic ) ) | |||||||||||||||||||||
3D reconstruction | Resolution: 6.9 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 8725 / Symmetry type: POINT | |||||||||||||||||||||
| Atomic model building | B value: 670.8 / Protocol: FLEXIBLE FIT / Space: REAL |
 Movie
Movie Controller
Controller










 PDBj
PDBj
