+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6roa | ||||||
|---|---|---|---|---|---|---|---|
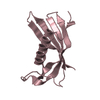
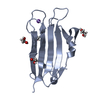
| Title | Crystal structure of V57G mutant of human cystatin C | ||||||
 Components Components | Cystatin-C | ||||||
 Keywords Keywords |  HYDROLASE / hydrolase inhibitor / monomeric structure HYDROLASE / hydrolase inhibitor / monomeric structure | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of collagen catabolic process / negative regulation of elastin catabolic process / negative regulation of blood vessel remodeling /  peptidase inhibitor activity / negative regulation of peptidase activity / negative regulation of extracellular matrix disassembly / peptidase inhibitor activity / negative regulation of peptidase activity / negative regulation of extracellular matrix disassembly /  regulation of tissue remodeling / cysteine-type endopeptidase inhibitor activity / regulation of tissue remodeling / cysteine-type endopeptidase inhibitor activity /  endopeptidase inhibitor activity / supramolecular fiber organization ...negative regulation of collagen catabolic process / negative regulation of elastin catabolic process / negative regulation of blood vessel remodeling / endopeptidase inhibitor activity / supramolecular fiber organization ...negative regulation of collagen catabolic process / negative regulation of elastin catabolic process / negative regulation of blood vessel remodeling /  peptidase inhibitor activity / negative regulation of peptidase activity / negative regulation of extracellular matrix disassembly / peptidase inhibitor activity / negative regulation of peptidase activity / negative regulation of extracellular matrix disassembly /  regulation of tissue remodeling / cysteine-type endopeptidase inhibitor activity / regulation of tissue remodeling / cysteine-type endopeptidase inhibitor activity /  endopeptidase inhibitor activity / supramolecular fiber organization / negative regulation of proteolysis / endopeptidase inhibitor activity / supramolecular fiber organization / negative regulation of proteolysis /  Post-translational protein phosphorylation / defense response / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / tertiary granule lumen / Post-translational protein phosphorylation / defense response / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / tertiary granule lumen /  amyloid-beta binding / amyloid-beta binding /  protease binding / vesicle / ficolin-1-rich granule lumen / Amyloid fiber formation / protease binding / vesicle / ficolin-1-rich granule lumen / Amyloid fiber formation /  endoplasmic reticulum lumen / Neutrophil degranulation / endoplasmic reticulum lumen / Neutrophil degranulation /  Golgi apparatus / Golgi apparatus /  endoplasmic reticulum / endoplasmic reticulum /  extracellular space / extracellular exosome / extracellular region / identical protein binding / extracellular space / extracellular exosome / extracellular region / identical protein binding /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.65 Å MOLECULAR REPLACEMENT / Resolution: 2.65 Å | ||||||
 Authors Authors | Orlikowska, M. / Behrendt, I. / Borek, D. / Otwinowski, Z. / Skowron, P. / Szymanska, A. | ||||||
 Citation Citation |  Journal: Febs J. / Year: 2020 Journal: Febs J. / Year: 2020Title: NMR and crystallographic structural studies of the extremely stable monomeric variant of human cystatin C with single amino acid substitution. Authors: Maszota-Zieleniak, M. / Jurczak, P. / Orlikowska, M. / Zhukov, I. / Borek, D. / Otwinowski, Z. / Skowron, P. / Pietralik, Z. / Kozak, M. / Szymanska, A. / Rodziewicz-Motowidlo, S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6roa.cif.gz 6roa.cif.gz | 104.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6roa.ent.gz pdb6roa.ent.gz | 81.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6roa.json.gz 6roa.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ro/6roa https://data.pdbj.org/pub/pdb/validation_reports/ro/6roa ftp://data.pdbj.org/pub/pdb/validation_reports/ro/6roa ftp://data.pdbj.org/pub/pdb/validation_reports/ro/6roa | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6rpvC  3nx0S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 13323.062 Da / Num. of mol.: 2 / Mutation: V57G Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: CST3 / Plasmid: pHD313 / Production host: Homo sapiens (human) / Gene: CST3 / Plasmid: pHD313 / Production host:   Escherichia coli (E. coli) / Variant (production host): C41(DE3) / References: UniProt: P01034 Escherichia coli (E. coli) / Variant (production host): C41(DE3) / References: UniProt: P01034#2: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.04 Å3/Da / Density % sol: 59.57 % |
|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: 0.2 M Na acetate, 0.1M Na cacodylate pH 6.5, 30% PEG 8000. |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 19-ID / Wavelength: 0.97921 Å / Beamline: 19-ID / Wavelength: 0.97921 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Jul 7, 2010 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.97921 Å / Relative weight: 1 : 0.97921 Å / Relative weight: 1 |
| Reflection | Resolution: 2.65→50 Å / Num. obs: 9095 / % possible obs: 96.8 % / Redundancy: 4.7 % / Biso Wilson estimate: 47.82 Å2 / Rmerge(I) obs: 0.127 / Net I/σ(I): 13.7 |
| Reflection shell | Resolution: 2.65→2.7 Å / Redundancy: 4.3 % / Rmerge(I) obs: 0.565 / Mean I/σ(I) obs: 2.05 / Num. unique obs: 9095 / % possible all: 96.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3NX0 Resolution: 2.65→50 Å / Cor.coef. Fo:Fc: 0.958 / Cor.coef. Fo:Fc free: 0.908 / SU B: 25.648 / SU ML: 0.249 / Cross valid method: THROUGHOUT / ESU R: 0.528 / ESU R Free: 0.314 / Details: HYDROGENS HAVE BEEN USED IF PRESENT IN THE INPUT
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 49.82 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.65→50 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj
