+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6jvx | ||||||
|---|---|---|---|---|---|---|---|
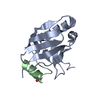
| Title | Crystal structure of RBM38 in complex with RNA | ||||||
 Components Components |
| ||||||
 Keywords Keywords | RNA BINDING PROTEIN/RNA / RBM38 /  RNA binding / RNA binding /  Translational regulation / Translational regulation /  RNA BINDING PROTEIN / RNA BINDING PROTEIN-RNA complex RNA BINDING PROTEIN / RNA BINDING PROTEIN-RNA complex | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of myotube differentiation / DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator / 3'-UTR-mediated mRNA stabilization /  regulation of RNA splicing / DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest / regulation of RNA splicing / DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest /  RNA splicing / mRNA 3'-UTR binding / RNA splicing / mRNA 3'-UTR binding /  mRNA processing / mRNA processing /  cell differentiation / cell differentiation /  regulation of cell cycle ...regulation of myotube differentiation / DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator / 3'-UTR-mediated mRNA stabilization / regulation of cell cycle ...regulation of myotube differentiation / DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator / 3'-UTR-mediated mRNA stabilization /  regulation of RNA splicing / DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest / regulation of RNA splicing / DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest /  RNA splicing / mRNA 3'-UTR binding / RNA splicing / mRNA 3'-UTR binding /  mRNA processing / mRNA processing /  cell differentiation / cell differentiation /  regulation of cell cycle / regulation of cell cycle /  cell cycle / negative regulation of cell population proliferation / cell cycle / negative regulation of cell population proliferation /  mRNA binding / mRNA binding /  RNA binding / RNA binding /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.301 Å MOLECULAR REPLACEMENT / Resolution: 2.301 Å | ||||||
 Authors Authors | Qian, K. / Li, M. / Wang, J. / Zhang, M. / Wang, M. | ||||||
 Citation Citation |  Journal: Biochem.J. / Year: 2020 Journal: Biochem.J. / Year: 2020Title: Structural basis for mRNA recognition by human RBM38. Authors: Qian, K. / Li, M. / Wang, J. / Zhang, M. / Wang, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6jvx.cif.gz 6jvx.cif.gz | 58.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6jvx.ent.gz pdb6jvx.ent.gz | 39.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6jvx.json.gz 6jvx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jv/6jvx https://data.pdbj.org/pub/pdb/validation_reports/jv/6jvx ftp://data.pdbj.org/pub/pdb/validation_reports/jv/6jvx ftp://data.pdbj.org/pub/pdb/validation_reports/jv/6jvx | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6jvyC  2cqdS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 13853.546 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: RBM38, RNPC1, SEB4 / Production host: Homo sapiens (human) / Gene: RBM38, RNPC1, SEB4 / Production host:   Escherichia coli (E. coli) / References: UniProt: Q9H0Z9 Escherichia coli (E. coli) / References: UniProt: Q9H0Z9 | ||
|---|---|---|---|
| #2: RNA chain | Mass: 3863.272 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)   Homo sapiens (human) Homo sapiens (human) | ||
| #3: Chemical |  Sulfate Sulfate#4: Water | ChemComp-HOH / |  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.39 Å3/Da / Density % sol: 63.71 % |
|---|---|
Crystal grow | Temperature: 289 K / Method: vapor diffusion, sitting drop / pH: 8 / Details: 0.1M Tris-HCl (pH8.0), 2M ammonium sulfate |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRF SSRF  / Beamline: BL17U / Wavelength: 0.9791 Å / Beamline: BL17U / Wavelength: 0.9791 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: DECTRIS EIGER X 16M / Detector: PIXEL / Date: Jul 15, 2018 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength : 0.9791 Å / Relative weight: 1 : 0.9791 Å / Relative weight: 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.3→50 Å / Num. obs: 10894 / % possible obs: 98.9 % / Redundancy: 4.9 % / Rmerge(I) obs: 0.101 / Rpim(I) all: 0.049 / Rrim(I) all: 0.113 / Χ2: 1.049 / Net I/σ(I): 9.8 / Num. measured all: 53598 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2cqd Resolution: 2.301→39.405 Å / SU ML: 0.33 / Cross valid method: THROUGHOUT / σ(F): 1.38 / Phase error: 23.37
| ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 128.63 Å2 / Biso mean: 43.1122 Å2 / Biso min: 15.44 Å2 | ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.301→39.405 Å
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Rfactor Rfree error: 0 / Total num. of bins used: 4
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: -30.1613 Å / Origin y: 25.7511 Å / Origin z: 4.679 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: (chain 'A' and resid 29 through 115) |
 Movie
Movie Controller
Controller












 PDBj
PDBj
































