[English] 日本語
 Yorodumi
Yorodumi- PDB-5z55: NMR Solution Structure of the Kringle domain of human receptor ty... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5z55 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | NMR Solution Structure of the Kringle domain of human receptor tyrosine kinase-like orphan receptor 1 (ROR1) | ||||||||||||||||||
 Components Components | Inactive tyrosine-protein kinase transmembrane receptor ROR1 | ||||||||||||||||||
 Keywords Keywords |  ONCOPROTEIN / ONCOPROTEIN /  ROR1 / ROR1 /  Kringle Kringle | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationWNT5A-dependent internalization of FZD2, FZD5 and ROR2 /  Wnt receptor activity / Wnt-protein binding / astrocyte development / inner ear development / Wnt receptor activity / Wnt-protein binding / astrocyte development / inner ear development /  coreceptor activity / axon terminus / coreceptor activity / axon terminus /  stress fiber / stress fiber /  transmembrane receptor protein tyrosine kinase activity / sensory perception of sound ...WNT5A-dependent internalization of FZD2, FZD5 and ROR2 / transmembrane receptor protein tyrosine kinase activity / sensory perception of sound ...WNT5A-dependent internalization of FZD2, FZD5 and ROR2 /  Wnt receptor activity / Wnt-protein binding / astrocyte development / inner ear development / Wnt receptor activity / Wnt-protein binding / astrocyte development / inner ear development /  coreceptor activity / axon terminus / coreceptor activity / axon terminus /  stress fiber / stress fiber /  transmembrane receptor protein tyrosine kinase activity / sensory perception of sound / transmembrane receptor protein tyrosine kinase activity / sensory perception of sound /  cell surface receptor protein tyrosine kinase signaling pathway / positive regulation of neuron projection development / positive regulation of NF-kappaB transcription factor activity / positive regulation of canonical NF-kappaB signal transduction / cell surface receptor protein tyrosine kinase signaling pathway / positive regulation of neuron projection development / positive regulation of NF-kappaB transcription factor activity / positive regulation of canonical NF-kappaB signal transduction /  receptor complex / receptor complex /  phosphorylation / phosphorylation /  cell surface / cell surface /  ATP binding / ATP binding /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | ||||||||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
| Method |  SOLUTION NMR / torsion angle dynamics SOLUTION NMR / torsion angle dynamics | ||||||||||||||||||
 Authors Authors | Ma, X. / Hu, K.F. | ||||||||||||||||||
| Funding support |  China, 5items China, 5items
| ||||||||||||||||||
 Citation Citation |  Journal: Biomol NMR Assign / Year: 2018 Journal: Biomol NMR Assign / Year: 2018Title: Backbone and side-chain chemical shift assignments of the kringle domain of human receptor tyrosine kinase-like orphan receptor 1 (ROR1). Authors: Ma, X. / Zhang, Y. / Liu, B. / Yang, J. / Hu, K. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5z55.cif.gz 5z55.cif.gz | 247.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5z55.ent.gz pdb5z55.ent.gz | 210 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5z55.json.gz 5z55.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/z5/5z55 https://data.pdbj.org/pub/pdb/validation_reports/z5/5z55 ftp://data.pdbj.org/pub/pdb/validation_reports/z5/5z55 ftp://data.pdbj.org/pub/pdb/validation_reports/z5/5z55 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 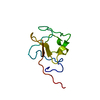
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 9562.522 Da / Num. of mol.: 1 / Fragment: Kringle domain Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: ROR1, NTRKR1 / Production host: Homo sapiens (human) / Gene: ROR1, NTRKR1 / Production host:   Escherichia coli (E. coli) / References: UniProt: Q01973 Escherichia coli (E. coli) / References: UniProt: Q01973 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Type: solution / Contents: 0.8 mM 99%U-13C, U-15N ROR1Kr, 94% H2O/6% D2O Details: Kringle domain of human receptor tyrosine kinase-like orphan receptor 1 (ROR1) Label: 13C_15N_ROR1Kr / Solvent system: 94% H2O/6% D2O |
|---|---|
| Sample | Conc.: 0.8 mM / Component: ROR1Kr / Isotopic labeling: 99%U-13C, U-15N |
| Sample conditions | Details: buffer: 25 mM Tris, 150 mM NaCl, 6% D2O and 0.02% NaN3 (w/v) at pH 8.0 and 298 K Ionic strength: 150 mM / Label: conditions_1 / pH: 8.0 / Pressure: 1 atm / Temperature: 298 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: torsion angle dynamics / Software ordinal: 6 | |||||||||||||||||||||
| NMR representative | Selection criteria: medoid | |||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 96 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller











 PDBj
PDBj




