+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5w0h | ||||||
|---|---|---|---|---|---|---|---|
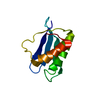
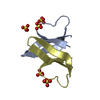
| Title | Structure of U2AF65 (U2AF2) RRM2 at 1.11 Angstrom Resolution | ||||||
 Components Components | Splicing factor U2AF 65 kDa subunit | ||||||
 Keywords Keywords | RNA BINDING PROTEIN/RNA / RNA SPLICING FACTOR /  RNA RECOGNITION MOTIF / RNA RECOGNITION MOTIF /  POLYPYRIMIDINE TRACT / POLYPYRIMIDINE TRACT /  RNA BINDING PROTEIN / RNA BINDING PROTEIN-RNA complex RNA BINDING PROTEIN / RNA BINDING PROTEIN-RNA complex | ||||||
| Function / homology |  Function and homology information Function and homology informationU2AF complex / poly-pyrimidine tract binding / pre-mRNA 3'-splice site binding / mRNA 3'-end processing / C2H2 zinc finger domain binding / commitment complex / Transport of Mature mRNA derived from an Intron-Containing Transcript / RNA Polymerase II Transcription Termination / U2-type prespliceosome / molecular function inhibitor activity ...U2AF complex / poly-pyrimidine tract binding / pre-mRNA 3'-splice site binding / mRNA 3'-end processing / C2H2 zinc finger domain binding / commitment complex / Transport of Mature mRNA derived from an Intron-Containing Transcript / RNA Polymerase II Transcription Termination / U2-type prespliceosome / molecular function inhibitor activity /  Protein hydroxylation / negative regulation of mRNA splicing, via spliceosome / negative regulation of protein ubiquitination / mRNA Splicing - Major Pathway / positive regulation of RNA splicing / Protein hydroxylation / negative regulation of mRNA splicing, via spliceosome / negative regulation of protein ubiquitination / mRNA Splicing - Major Pathway / positive regulation of RNA splicing /  spliceosomal complex / spliceosomal complex /  mRNA processing / mRNA processing /  mRNA splicing, via spliceosome / nuclear speck / mRNA splicing, via spliceosome / nuclear speck /  enzyme binding / enzyme binding /  RNA binding / RNA binding /  nucleoplasm / nucleoplasm /  nucleus nucleusSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.11 Å molecular replacement / Resolution: 1.11 Å | ||||||
 Authors Authors | Agrawal, A.A. / Jenkins, J.L. / Kielkopf, C.L. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2017 Journal: Biochemistry / Year: 2017Title: Cancer-Associated Mutations Mapped on High-Resolution Structures of the U2AF2 RNA Recognition Motifs. Authors: Glasser, E. / Agrawal, A.A. / Jenkins, J.L. / Kielkopf, C.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5w0h.cif.gz 5w0h.cif.gz | 71.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5w0h.ent.gz pdb5w0h.ent.gz | 53.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5w0h.json.gz 5w0h.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/w0/5w0h https://data.pdbj.org/pub/pdb/validation_reports/w0/5w0h ftp://data.pdbj.org/pub/pdb/validation_reports/w0/5w0h ftp://data.pdbj.org/pub/pdb/validation_reports/w0/5w0h | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5w0gC  2g4bS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 8966.237 Da / Num. of mol.: 1 / Fragment: RNA Recognition Motif 2 (RRM2), residues 258-336 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: U2AF2, U2AF65 / Plasmid: PGEX-6P / Production host: Homo sapiens (human) / Gene: U2AF2, U2AF65 / Plasmid: PGEX-6P / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21 / References: UniProt: P26368 Escherichia coli (E. coli) / Strain (production host): BL21 / References: UniProt: P26368 |
|---|---|
| #2: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.59 Å3/Da / Density % sol: 52.5 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 5.5 Details: 26% (w/v) PEG 8000, 0.1 M NaOAc, 0.2 mM TCEP, 10% (v/v) dioxane, 0.2 ul BigCHAP, 0.05 M potassium phosphate monobasic |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CHESS CHESS  / Beamline: F1 / Wavelength: 0.9779 Å / Beamline: F1 / Wavelength: 0.9779 Å |
| Detector | Type: ADSC QUANTUM 270 / Detector: CCD / Date: May 22, 2013 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.9779 Å / Relative weight: 1 : 0.9779 Å / Relative weight: 1 |
| Reflection | Resolution: 1.11→20.69 Å / Num. obs: 36104 / % possible obs: 97.2 % / Redundancy: 5.8 % / Biso Wilson estimate: 11.44 Å2 / Rmerge(I) obs: 0.081 / Net I/σ(I): 27.8 |
| Reflection shell | Resolution: 1.11→1.13 Å / Redundancy: 3.7 % / Rmerge(I) obs: 0.278 / Mean I/σ(I) obs: 4.5 / % possible all: 92.2 |
-Phasing
Phasing | Method:  molecular replacement molecular replacement |
|---|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2G4B RRM2 Resolution: 1.11→20.69 Å / SU ML: 0.09 / Cross valid method: FREE R-VALUE / σ(F): 1.33 / Phase error: 15.32
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 67.74 Å2 / Biso mean: 18.5012 Å2 / Biso min: 7.31 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.11→20.69 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Rfactor Rfree error: 0 / Total num. of bins used: 14
|
 Movie
Movie Controller
Controller











 PDBj
PDBj



