+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5o10 | ||||||
|---|---|---|---|---|---|---|---|
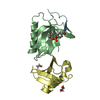
| Title | Y48H mutant of human cytochrome c | ||||||
 Components Components | Cytochrome c | ||||||
 Keywords Keywords |  ELECTRON TRANSFER / ELECTRON TRANSFER /  APOPTOSIS / APOPTOSIS /  Heme / Heme /  haem / haem /  cytochrome c / cytochrome c /  metalloprotein metalloprotein | ||||||
| Function / homology |  Function and homology information Function and homology informationFormation of apoptosome /  apoptosome / activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c / Release of apoptotic factors from the mitochondria / Respiratory electron transport / apoptosome / activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c / Release of apoptotic factors from the mitochondria / Respiratory electron transport /  cellular respiration / Regulation of the apoptosome activity / Activation of caspases through apoptosome-mediated cleavage / SMAC (DIABLO) binds to IAPs / SMAC(DIABLO)-mediated dissociation of IAP:caspase complexes ...Formation of apoptosome / cellular respiration / Regulation of the apoptosome activity / Activation of caspases through apoptosome-mediated cleavage / SMAC (DIABLO) binds to IAPs / SMAC(DIABLO)-mediated dissociation of IAP:caspase complexes ...Formation of apoptosome /  apoptosome / activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c / Release of apoptotic factors from the mitochondria / Respiratory electron transport / apoptosome / activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c / Release of apoptotic factors from the mitochondria / Respiratory electron transport /  cellular respiration / Regulation of the apoptosome activity / Activation of caspases through apoptosome-mediated cleavage / SMAC (DIABLO) binds to IAPs / SMAC(DIABLO)-mediated dissociation of IAP:caspase complexes / mitochondrial electron transport, cytochrome c to oxygen / mitochondrial electron transport, ubiquinol to cytochrome c / Detoxification of Reactive Oxygen Species / cellular respiration / Regulation of the apoptosome activity / Activation of caspases through apoptosome-mediated cleavage / SMAC (DIABLO) binds to IAPs / SMAC(DIABLO)-mediated dissociation of IAP:caspase complexes / mitochondrial electron transport, cytochrome c to oxygen / mitochondrial electron transport, ubiquinol to cytochrome c / Detoxification of Reactive Oxygen Species /  Pyroptosis / Pyroptosis /  respirasome / intrinsic apoptotic signaling pathway / TP53 Regulates Metabolic Genes / Transcriptional activation of mitochondrial biogenesis / respirasome / intrinsic apoptotic signaling pathway / TP53 Regulates Metabolic Genes / Transcriptional activation of mitochondrial biogenesis /  mitochondrial intermembrane space / Cytoprotection by HMOX1 / mitochondrial intermembrane space / Cytoprotection by HMOX1 /  mitochondrial inner membrane / mitochondrial inner membrane /  electron transfer activity / electron transfer activity /  heme binding / heme binding /  mitochondrion / mitochondrion /  metal ion binding / metal ion binding /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.36 Å MOLECULAR REPLACEMENT / Resolution: 1.36 Å | ||||||
 Authors Authors | Moreno-Chicano, T. / Deacon, O.M. / Hough, M.A. / Worrall, J.A.R. | ||||||
| Funding support |  United Kingdom, 1items United Kingdom, 1items
| ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2017 Journal: Biochemistry / Year: 2017Title: Heightened Dynamics of the Oxidized Y48H Variant of Human Cytochrome c Increases Its Peroxidatic Activity. Authors: Deacon, O.M. / Karsisiotis, A.I. / Moreno-Chicano, T. / Hough, M.A. / Macdonald, C. / Blumenschein, T.M.A. / Wilson, M.T. / Moore, G.R. / Worrall, J.A.R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5o10.cif.gz 5o10.cif.gz | 109.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5o10.ent.gz pdb5o10.ent.gz | 83.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5o10.json.gz 5o10.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/o1/5o10 https://data.pdbj.org/pub/pdb/validation_reports/o1/5o10 ftp://data.pdbj.org/pub/pdb/validation_reports/o1/5o10 ftp://data.pdbj.org/pub/pdb/validation_reports/o1/5o10 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3zcfS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  Mass: 11615.556 Da / Num. of mol.: 2 / Mutation: Y48H Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: CYCS, CYC Homo sapiens (human) / Gene: CYCS, CYCProduction host:   Escherichia coli 'BL21-Gold(DE3)pLysS AG' (bacteria) Escherichia coli 'BL21-Gold(DE3)pLysS AG' (bacteria)References: UniProt: P99999 #2: Chemical |  Heme C Heme C#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.43 Å3/Da / Density % sol: 49.31 % |
|---|---|
Crystal grow | Temperature: 291 K / Method: vapor diffusion, sitting drop / pH: 7.5 / Details: 33% PEG 6000 100mM Tris pH 7.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X10SA / Wavelength: 0.999 Å / Beamline: X10SA / Wavelength: 0.999 Å |
| Detector | Type: DECTRIS PILATUS 6M-F / Detector: PIXEL / Date: Nov 11, 2016 / Details: Mirrors |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.999 Å / Relative weight: 1 : 0.999 Å / Relative weight: 1 |
| Reflection | Resolution: 1.36→54.25 Å / Num. obs: 45812 / % possible obs: 95.3 % / Redundancy: 6.2 % / CC1/2: 0.998 / Rmerge(I) obs: 0.072 / Rpim(I) all: 0.031 / Net I/av σ(I): 13.3 / Net I/σ(I): 13.3 |
| Reflection shell | Resolution: 1.36→1.39 Å / Redundancy: 5.4 % / Rmerge(I) obs: 1.115 / Mean I/σ(I) obs: 1.4 / Num. unique obs: 2124 / CC1/2: 0.51 / Rpim(I) all: 0.51 / % possible all: 90.2 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3zcf Resolution: 1.36→54.25 Å / Cor.coef. Fo:Fc: 0.974 / Cor.coef. Fo:Fc free: 0.97 / SU B: 1.963 / SU ML: 0.035 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.056 / ESU R Free: 0.052 Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 63.31 Å2 / Biso mean: 18.567 Å2 / Biso min: 8.26 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.36→54.25 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.362→1.397 Å / Rfactor Rfree error: 0 / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller












 PDBj
PDBj























