[English] 日本語
 Yorodumi
Yorodumi- PDB-5j4o: Structure of human erythrocytic Spectrin alpha chain repeats 16-17 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5j4o | ||||||
|---|---|---|---|---|---|---|---|
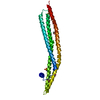
| Title | Structure of human erythrocytic Spectrin alpha chain repeats 16-17 | ||||||
 Components Components | Spectrin alpha chain, erythrocytic 1 | ||||||
 Keywords Keywords |  STRUCTURAL PROTEIN / STRUCTURAL PROTEIN /  erythrocyte / erythrocyte /  spectrin / spectrin /  helical bundle / helical bundle /  domains domains | ||||||
| Function / homology |  Function and homology information Function and homology information cuticular plate / cuticular plate /  spectrin / lymphocyte homeostasis / spectrin-associated cytoskeleton / porphyrin-containing compound biosynthetic process / plasma membrane organization / actin filament capping / Interaction between L1 and Ankyrins / cortical actin cytoskeleton / spectrin / lymphocyte homeostasis / spectrin-associated cytoskeleton / porphyrin-containing compound biosynthetic process / plasma membrane organization / actin filament capping / Interaction between L1 and Ankyrins / cortical actin cytoskeleton /  hemopoiesis ... hemopoiesis ... cuticular plate / cuticular plate /  spectrin / lymphocyte homeostasis / spectrin-associated cytoskeleton / porphyrin-containing compound biosynthetic process / plasma membrane organization / actin filament capping / Interaction between L1 and Ankyrins / cortical actin cytoskeleton / spectrin / lymphocyte homeostasis / spectrin-associated cytoskeleton / porphyrin-containing compound biosynthetic process / plasma membrane organization / actin filament capping / Interaction between L1 and Ankyrins / cortical actin cytoskeleton /  hemopoiesis / COPI-mediated anterograde transport / positive regulation of T cell proliferation / NCAM signaling for neurite out-growth / actin filament organization / cell projection / cytoplasmic side of plasma membrane / structural constituent of cytoskeleton / hemopoiesis / COPI-mediated anterograde transport / positive regulation of T cell proliferation / NCAM signaling for neurite out-growth / actin filament organization / cell projection / cytoplasmic side of plasma membrane / structural constituent of cytoskeleton /  actin filament binding / actin filament binding /  actin cytoskeleton / actin cytoskeleton /  cell junction / regulation of cell shape / actin cytoskeleton organization / RAF/MAP kinase cascade / cell junction / regulation of cell shape / actin cytoskeleton organization / RAF/MAP kinase cascade /  axon / axon /  calcium ion binding / calcium ion binding /  plasma membrane / plasma membrane /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.54 Å MOLECULAR REPLACEMENT / Resolution: 1.54 Å | ||||||
 Authors Authors | Cutts, E.E. / Vakonakis, I. | ||||||
| Funding support |  United Kingdom, 1items United Kingdom, 1items
| ||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Interactions of Plasmodium falciparum KAHRP and PfEMP1 with the host cytoskeleton suggest a model for cytoadherent protrusions on the infected erythrocyte surface Authors: Cutts, E.E. / Laasch, N. / Reiter, D.M. / Trenker, R. / Slater, L.M. / Stansfeld, P.J. / Vakonakis, I. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5j4o.cif.gz 5j4o.cif.gz | 200.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5j4o.ent.gz pdb5j4o.ent.gz | 161.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5j4o.json.gz 5j4o.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/j4/5j4o https://data.pdbj.org/pub/pdb/validation_reports/j4/5j4o ftp://data.pdbj.org/pub/pdb/validation_reports/j4/5j4o ftp://data.pdbj.org/pub/pdb/validation_reports/j4/5j4o | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1u5pS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 26757.439 Da / Num. of mol.: 1 / Fragment: UNP residues 1599-1826 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Tissue: Blood Homo sapiens (human) / Tissue: Blood / Cell: Erythrocytes / Gene: SPTA1, SPTA / Plasmid: pET16c / Cell: Erythrocytes / Gene: SPTA1, SPTA / Plasmid: pET16cDetails (production host): Modified to include 3C protease site Production host:   Escherichia coli BL21(DE3) (bacteria) / Variant (production host): Rosetta2 / References: UniProt: P02549 Escherichia coli BL21(DE3) (bacteria) / Variant (production host): Rosetta2 / References: UniProt: P02549 | ||||
|---|---|---|---|---|---|
| #2: Chemical | ChemComp-EDO /  Ethylene glycol Ethylene glycol#3: Chemical |  Polyethylene glycol Polyethylene glycol#4: Water | ChemComp-HOH / |  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.37 Å3/Da / Density % sol: 48.09 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, sitting drop / pH: 6.5 Details: Molecular Dimensions Morpheus crystallisation screen, condition A2: 0.03M MgCl2, 0.03M CaCl2, 0.1M MES/Imidazole pH 6.5, 20% v/v Ethylene glycol, 10% v/v PEG 8000. |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I04-1 / Wavelength: 0.91741 Å / Beamline: I04-1 / Wavelength: 0.91741 Å |
| Detector | Type: DECTRIS PILATUS 2M / Detector: PIXEL / Date: Feb 15, 2015 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.91741 Å / Relative weight: 1 : 0.91741 Å / Relative weight: 1 |
| Reflection | Resolution: 1.54→38.57 Å / Num. obs: 38551 / % possible obs: 99.6 % / Redundancy: 6.4 % / Biso Wilson estimate: 23.5 Å2 / CC1/2: 0.998 / Rmerge(I) obs: 0.065 / Net I/σ(I): 13.1 |
| Reflection shell | Resolution: 1.54→1.58 Å / Redundancy: 6.2 % / Rmerge(I) obs: 0.926 / Mean I/σ(I) obs: 1.7 / % possible all: 99.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1U5P Resolution: 1.54→30.18 Å / Cor.coef. Fo:Fc: 0.9362 / Cor.coef. Fo:Fc free: 0.925 / SU R Cruickshank DPI: 0.101 / Cross valid method: THROUGHOUT / σ(F): 0 / SU R Blow DPI: 0.089 / SU Rfree Blow DPI: 0.085 / SU Rfree Cruickshank DPI: 0.084
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 35.6 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.207 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: 1 / Resolution: 1.54→30.18 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.54→1.58 Å / Total num. of bins used: 19
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: -3.0795 Å / Origin y: 5.4751 Å / Origin z: -23.0408 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: { A|* } |
 Movie
Movie Controller
Controller










 PDBj
PDBj












