+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5dhe | ||||||
|---|---|---|---|---|---|---|---|
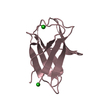
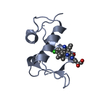
| Title | Crystal structure of ChBD3 from Thermococcus kodakarensis KOD1 | ||||||
 Components Components | Chitinase | ||||||
 Keywords Keywords |  HYDROLASE / HYDROLASE /  chitin / chitin /  chitinase / chitin binding domain chitinase / chitin binding domain | ||||||
| Function / homology |  Function and homology information Function and homology information polysaccharide binding / polysaccharide binding /  chitin binding / hydrolase activity, hydrolyzing O-glycosyl compounds / carbohydrate metabolic process / extracellular region chitin binding / hydrolase activity, hydrolyzing O-glycosyl compounds / carbohydrate metabolic process / extracellular regionSimilarity search - Function | ||||||
| Biological species |    Thermococcus kodakarensis KOD1 (archaea) Thermococcus kodakarensis KOD1 (archaea) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 1.6 Å SAD / Resolution: 1.6 Å | ||||||
 Authors Authors | Niwa, S. / Hibi, M. / Takeda, K. / Miki, K. | ||||||
| Funding support |  Japan, 1items Japan, 1items
| ||||||
 Citation Citation |  Journal: Febs Lett. / Year: 2016 Journal: Febs Lett. / Year: 2016Title: Crystal structures of chitin binding domains of chitinase from Thermococcus kodakarensis KOD1 Authors: Hanazono, Y. / Takeda, K. / Niwa, S. / Hibi, M. / Takahashi, N. / Kanai, T. / Atomi, H. / Miki, K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5dhe.cif.gz 5dhe.cif.gz | 59 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5dhe.ent.gz pdb5dhe.ent.gz | 42.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5dhe.json.gz 5dhe.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dh/5dhe https://data.pdbj.org/pub/pdb/validation_reports/dh/5dhe ftp://data.pdbj.org/pub/pdb/validation_reports/dh/5dhe ftp://data.pdbj.org/pub/pdb/validation_reports/dh/5dhe | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  / Chitinase / containing dual catalytic domains / Chitinase / containing dual catalytic domainsMass: 11008.195 Da / Num. of mol.: 2 / Fragment: ChBD3 domain, UNP residues 764-863 Source method: isolated from a genetically manipulated source Source: (gene. exp.)    Thermococcus kodakarensis KOD1 (archaea) Thermococcus kodakarensis KOD1 (archaea)Strain: KOD1 / Gene: Pk-chiA, TK1765 / Production host:   Escherichia coli (E. coli) / References: UniProt: Q9UWR7, Escherichia coli (E. coli) / References: UniProt: Q9UWR7,  chitinase chitinase#2: Chemical | ChemComp-GOL /  Glycerol Glycerol#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.34 Å3/Da / Density % sol: 47.43 % |
|---|---|
Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop / pH: 6.5 / Details: sodium citrate, glycerol, MES, SDS |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SPring-8 SPring-8  / Beamline: BL26B2 / Wavelength: 0.9 Å / Beamline: BL26B2 / Wavelength: 0.9 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: Dec 9, 2013 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength : 0.9 Å / Relative weight: 1 : 0.9 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Redundancy: 4 % / Number: 123739 / Rmerge(I) obs: 0.085 / Χ2: 1.6 / D res high: 2.4 Å / D res low: 50 Å / Num. obs: 31247 / % possible obs: 98.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diffraction reflection shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.6→30 Å / Num. obs: 26632 / % possible obs: 98.9 % / Redundancy: 4 % / Rmerge(I) obs: 0.061 / Χ2: 1.869 / Net I/av σ(I): 28.806 / Net I/σ(I): 13.6 / Num. measured all: 107174 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1 / Rejects: 0
|
-Phasing
Phasing | Method:  SAD SAD | |||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Phasing MAD | D res high: 3 Å / D res low: 15 Å / FOM : 0.39 / Reflection: 7742 | |||||||||||||||||||||||||||||||||||
| Phasing MAD set site |
| |||||||||||||||||||||||||||||||||||
| Phasing MAD shell |
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  SAD / Resolution: 1.6→30 Å / FOM work R set: 0.8763 / Cross valid method: FREE R-VALUE / σ(F): 0 SAD / Resolution: 1.6→30 Å / FOM work R set: 0.8763 / Cross valid method: FREE R-VALUE / σ(F): 0
| ||||||||||||||||||||||||
| Solvent computation | Bsol: 44.2818 Å2 | ||||||||||||||||||||||||
| Displacement parameters | Biso max: 51.52 Å2 / Biso mean: 14.7357 Å2 / Biso min: 1.95 Å2
| ||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.6→30 Å
| ||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj



