+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4u7i | ||||||
|---|---|---|---|---|---|---|---|
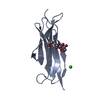
| Title | Structure of the complex of Spartin MIT and IST1 MIM | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  PROTEIN TRANSPORT / PROTEIN TRANSPORT /  Complex / MIM3 Complex / MIM3 | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of collateral sprouting in absence of injury / viral capsid secondary envelopment / MIT domain binding /  abscission / ESCRT III complex disassembly / cytoskeleton-dependent cytokinesis / collateral sprouting / Sealing of the nuclear envelope (NE) by ESCRT-III / positive regulation of collateral sprouting / lipid droplet organization ...negative regulation of collateral sprouting in absence of injury / viral capsid secondary envelopment / MIT domain binding / abscission / ESCRT III complex disassembly / cytoskeleton-dependent cytokinesis / collateral sprouting / Sealing of the nuclear envelope (NE) by ESCRT-III / positive regulation of collateral sprouting / lipid droplet organization ...negative regulation of collateral sprouting in absence of injury / viral capsid secondary envelopment / MIT domain binding /  abscission / ESCRT III complex disassembly / cytoskeleton-dependent cytokinesis / collateral sprouting / Sealing of the nuclear envelope (NE) by ESCRT-III / positive regulation of collateral sprouting / lipid droplet organization / abscission / ESCRT III complex disassembly / cytoskeleton-dependent cytokinesis / collateral sprouting / Sealing of the nuclear envelope (NE) by ESCRT-III / positive regulation of collateral sprouting / lipid droplet organization /  multivesicular body assembly / collateral sprouting in absence of injury / Flemming body / neuromuscular process / positive regulation of proteolysis / negative regulation of BMP signaling pathway / viral release from host cell / multivesicular body assembly / collateral sprouting in absence of injury / Flemming body / neuromuscular process / positive regulation of proteolysis / negative regulation of BMP signaling pathway / viral release from host cell /  endoplasmic reticulum-Golgi intermediate compartment / adipose tissue development / BMP signaling pathway / endoplasmic reticulum-Golgi intermediate compartment / adipose tissue development / BMP signaling pathway /  lipid droplet / regulation of mitochondrial membrane potential / establishment of protein localization / lipid droplet / regulation of mitochondrial membrane potential / establishment of protein localization /  protein localization / azurophil granule lumen / protein localization / azurophil granule lumen /  protein transport / protein transport /  nuclear envelope / midbody / mitochondrial outer membrane / nuclear envelope / midbody / mitochondrial outer membrane /  cadherin binding / protein domain specific binding / cadherin binding / protein domain specific binding /  cell division / intracellular membrane-bounded organelle / cell division / intracellular membrane-bounded organelle /  centrosome / centrosome /  synapse / synapse /  ubiquitin protein ligase binding / ubiquitin protein ligase binding /  chromatin / Neutrophil degranulation / protein-containing complex binding / extracellular exosome / extracellular region / identical protein binding / chromatin / Neutrophil degranulation / protein-containing complex binding / extracellular exosome / extracellular region / identical protein binding /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.794 Å MOLECULAR REPLACEMENT / Resolution: 1.794 Å | ||||||
 Authors Authors | Guo, E.Z. / Xu, Z. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2015 Journal: J.Biol.Chem. / Year: 2015Title: Distinct Mechanisms of Recognizing Endosomal Sorting Complex Required for Transport III (ESCRT-III) Protein IST1 by Different Microtubule Interacting and Trafficking (MIT) Domains. Authors: Guo, E.Z. / Xu, Z. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4u7i.cif.gz 4u7i.cif.gz | 62.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4u7i.ent.gz pdb4u7i.ent.gz | 44.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4u7i.json.gz 4u7i.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/u7/4u7i https://data.pdbj.org/pub/pdb/validation_reports/u7/4u7i ftp://data.pdbj.org/pub/pdb/validation_reports/u7/4u7i ftp://data.pdbj.org/pub/pdb/validation_reports/u7/4u7i | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4u7eC  4u7yC  2dl1S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 10794.293 Da / Num. of mol.: 1 / Fragment: MIT domain (UNP residues 8-101) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: SPG20, KIAA0610, TAHCCP1 / Production host: Homo sapiens (human) / Gene: SPG20, KIAA0610, TAHCCP1 / Production host:   Escherichia coli (E. coli) / Strain (production host): Rosetta (DE3) / References: UniProt: Q8N0X7 Escherichia coli (E. coli) / Strain (production host): Rosetta (DE3) / References: UniProt: Q8N0X7 |
|---|---|
| #2: Protein/peptide | Mass: 3009.262 Da / Num. of mol.: 1 / Fragment: UNP residues 341-364 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: IST1, KIAA0174 / Production host: Homo sapiens (human) / Gene: IST1, KIAA0174 / Production host:   Escherichia coli (E. coli) / Strain (production host): Rosetta (DE3) / References: UniProt: P53990 Escherichia coli (E. coli) / Strain (production host): Rosetta (DE3) / References: UniProt: P53990 |
| #3: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.24 Å3/Da / Density % sol: 45.1 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, sitting drop / pH: 7.3 / Details: 2.9 M Na-malonate / PH range: 6 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 21-ID-F / Wavelength: 0.97828 Å / Beamline: 21-ID-F / Wavelength: 0.97828 Å |
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: Aug 9, 2013 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.97828 Å / Relative weight: 1 : 0.97828 Å / Relative weight: 1 |
| Reflection | Resolution: 1.79→29.61 Å / Num. obs: 12400 / % possible obs: 99.6 % / Redundancy: 8.1 % / Rmerge(I) obs: 0.081 / Net I/σ(I): 43 |
| Reflection shell | Redundancy: 8.4 % / Rmerge(I) obs: 0.584 / Mean I/σ(I) obs: 4 / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2DL1 Resolution: 1.794→29.61 Å / SU ML: 0.19 / Cross valid method: NONE / σ(F): 1.35 / Phase error: 23.94 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.794→29.61 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: -12.3844 Å / Origin y: 6.1248 Å / Origin z: -7.1335 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: all |
 Movie
Movie Controller
Controller












 PDBj
PDBj
