+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4rwh | ||||||
|---|---|---|---|---|---|---|---|
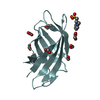
| Title | Crystal structure of T cell costimulatory ligand B7-1 (CD80) | ||||||
 Components Components | T-lymphocyte activation antigen CD80 | ||||||
 Keywords Keywords |  SIGNALING PROTEIN / Ig fold / T cell costimulatory ligand B7-1 SIGNALING PROTEIN / Ig fold / T cell costimulatory ligand B7-1 | ||||||
| Function / homology |  Function and homology information Function and homology informationCD28 dependent PI3K/Akt signaling / CD28 dependent Vav1 pathway / CD28 co-stimulation / CTLA4 inhibitory signaling / protein complex involved in cell adhesion / PIP3 activates AKT signaling / positive regulation of alpha-beta T cell proliferation / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling /  coreceptor activity / positive regulation of T cell proliferation ...CD28 dependent PI3K/Akt signaling / CD28 dependent Vav1 pathway / CD28 co-stimulation / CTLA4 inhibitory signaling / protein complex involved in cell adhesion / PIP3 activates AKT signaling / positive regulation of alpha-beta T cell proliferation / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling / coreceptor activity / positive regulation of T cell proliferation ...CD28 dependent PI3K/Akt signaling / CD28 dependent Vav1 pathway / CD28 co-stimulation / CTLA4 inhibitory signaling / protein complex involved in cell adhesion / PIP3 activates AKT signaling / positive regulation of alpha-beta T cell proliferation / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling /  coreceptor activity / positive regulation of T cell proliferation / negative regulation of T cell proliferation / T cell costimulation / positive regulation of peptidyl-tyrosine phosphorylation / cellular response to lipopolysaccharide / membrane => GO:0016020 / cell surface receptor signaling pathway / coreceptor activity / positive regulation of T cell proliferation / negative regulation of T cell proliferation / T cell costimulation / positive regulation of peptidyl-tyrosine phosphorylation / cellular response to lipopolysaccharide / membrane => GO:0016020 / cell surface receptor signaling pathway /  immune response / external side of plasma membrane / immune response / external side of plasma membrane /  cell surface / cell surface /  signal transduction signal transductionSimilarity search - Function | ||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.802 Å MOLECULAR REPLACEMENT / Resolution: 1.802 Å | ||||||
 Authors Authors | Fedorov, A.A. / Fedorov, E.V. / Samanta, D. / Hillerich, B. / Seidel, R.D. / Almo, S.C. | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Crystal structure of T cell costimulatory ligand B7-1 (CD80) Authors: Fedorov, A.A. / Fedorov, E.V. / Samanta, D. / Hillerich, B. / Seidel, R.D. / Almo, S.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4rwh.cif.gz 4rwh.cif.gz | 56.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4rwh.ent.gz pdb4rwh.ent.gz | 40.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4rwh.json.gz 4rwh.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/rw/4rwh https://data.pdbj.org/pub/pdb/validation_reports/rw/4rwh ftp://data.pdbj.org/pub/pdb/validation_reports/rw/4rwh ftp://data.pdbj.org/pub/pdb/validation_reports/rw/4rwh | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1dr9S S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 12335.271 Da / Num. of mol.: 1 / Fragment: UNP residues 44-144 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Gene: Cd80, B7 / Production host: Mus musculus (house mouse) / Gene: Cd80, B7 / Production host:   Escherichia coli (E. coli) / References: UniProt: Q00609 Escherichia coli (E. coli) / References: UniProt: Q00609 |
|---|---|
| #2: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.81 Å3/Da / Density % sol: 31.88 % |
|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 7 Details: 2.4M sodium malonate, pH 7.0, VAPOR DIFFUSION, SITTING DROP, temperature 293.0K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X29A / Wavelength: 1.075 Å / Beamline: X29A / Wavelength: 1.075 Å |
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Jul 3, 2014 |
| Radiation | Monochromator: Si 111 CHANNEL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.075 Å / Relative weight: 1 : 1.075 Å / Relative weight: 1 |
| Reflection | Resolution: 1.802→28.534 Å / Num. all: 11322 / Num. obs: 11322 / % possible obs: 99.46 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1DR9 Resolution: 1.802→28.534 Å / SU ML: 0.15 / σ(F): 0 / Phase error: 31.03 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.802→28.534 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 68.0483 Å / Origin y: 11.3089 Å / Origin z: 1.2083 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: chain A |
 Movie
Movie Controller
Controller












 PDBj
PDBj







