| Entry | Database: PDB / ID: 4b86
|
|---|
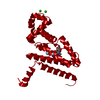
| Title | Crystal structure of the MSL1-MSL2 complex (3.5A) |
|---|
 Components Components | - MALE-SPECIFIC LETHAL 1 HOMOLOG
- MALE-SPECIFIC LETHAL 2 HOMOLOG
|
|---|
 Keywords Keywords |  GENE REGULATION / GENE REGULATION /  DOSAGE COMPENSATION / DOSAGE COMPENSATION /  CHROMATIN CHROMATIN |
|---|
| Function / homology |  Function and homology information Function and homology information |
|---|
| Biological species |   HOMO SAPIENS (human) HOMO SAPIENS (human) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 3.5 Å MAD / Resolution: 3.5 Å |
|---|
 Authors Authors | Hallacli, E. / Lipp, M. / Georgiev, P. / Spielman, C. / Cusack, S. / Akhtar, A. / Kadlec, J. |
|---|
 Citation Citation |  Journal: Mol.Cell / Year: 2012 Journal: Mol.Cell / Year: 2012
Title: Msl1-Mediated Dimerization of the Dosage Compensation Complex is Essential for Male X-Chromosome Regulation in Drosophila.
Authors: Hallacli, E. / Lipp, M. / Georgiev, P. / Spielman, C. / Cusack, S. / Akhtar, A. / Kadlec, J. |
|---|
| History | | Deposition | Aug 24, 2012 | Deposition site: PDBE / Processing site: PDBE |
|---|
| Revision 1.0 | Feb 6, 2013 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | May 8, 2024 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Other / Refinement description
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_struct_conn_angle / struct_conn / struct_ncs_dom_lim / struct_site
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_ncs_dom_lim.beg_auth_comp_id / _struct_ncs_dom_lim.beg_label_asym_id / _struct_ncs_dom_lim.beg_label_comp_id / _struct_ncs_dom_lim.beg_label_seq_id / _struct_ncs_dom_lim.end_auth_comp_id / _struct_ncs_dom_lim.end_label_asym_id / _struct_ncs_dom_lim.end_label_comp_id / _struct_ncs_dom_lim.end_label_seq_id / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords GENE REGULATION /
GENE REGULATION /  DOSAGE COMPENSATION /
DOSAGE COMPENSATION /  CHROMATIN
CHROMATIN Function and homology information
Function and homology information Transferases; Acyltransferases; Aminoacyltransferases /
Transferases; Acyltransferases; Aminoacyltransferases /  ubiquitin protein ligase activity / HATs acetylate histones / protein ubiquitination / nuclear speck /
ubiquitin protein ligase activity / HATs acetylate histones / protein ubiquitination / nuclear speck /  chromatin remodeling /
chromatin remodeling /  chromatin binding / positive regulation of DNA-templated transcription /
chromatin binding / positive regulation of DNA-templated transcription /  nucleoplasm ...MSL complex /
nucleoplasm ...MSL complex /  Transferases; Acyltransferases; Aminoacyltransferases /
Transferases; Acyltransferases; Aminoacyltransferases /  ubiquitin protein ligase activity / HATs acetylate histones / protein ubiquitination / nuclear speck /
ubiquitin protein ligase activity / HATs acetylate histones / protein ubiquitination / nuclear speck /  chromatin remodeling /
chromatin remodeling /  chromatin binding / positive regulation of DNA-templated transcription /
chromatin binding / positive regulation of DNA-templated transcription /  nucleoplasm /
nucleoplasm /  metal ion binding /
metal ion binding /  nucleus
nucleus
 HOMO SAPIENS (human)
HOMO SAPIENS (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MAD / Resolution: 3.5 Å
MAD / Resolution: 3.5 Å  Authors
Authors Citation
Citation Journal: Mol.Cell / Year: 2012
Journal: Mol.Cell / Year: 2012 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4b86.cif.gz
4b86.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4b86.ent.gz
pdb4b86.ent.gz PDB format
PDB format 4b86.json.gz
4b86.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/b8/4b86
https://data.pdbj.org/pub/pdb/validation_reports/b8/4b86 ftp://data.pdbj.org/pub/pdb/validation_reports/b8/4b86
ftp://data.pdbj.org/pub/pdb/validation_reports/b8/4b86 Links
Links Assembly
Assembly



 Movie
Movie Controller
Controller











 PDBj
PDBj