[English] 日本語
 Yorodumi
Yorodumi- PDB-4ayn: Structure of the C-terminal barrel of Neisseria meningitidis FHbp... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4ayn | ||||||
|---|---|---|---|---|---|---|---|
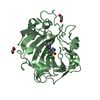
| Title | Structure of the C-terminal barrel of Neisseria meningitidis FHbp Variant 2 | ||||||
 Components Components | FACTOR H-BINDING PROTEIN | ||||||
 Keywords Keywords |  IMMUNE SYSTEM / IMMUNE SYSTEM /  ANTIGENS / ANTIGENS /  COMPLEMENT FACTOR H / VACCINES COMPLEMENT FACTOR H / VACCINES | ||||||
| Function / homology | Factor H binding protein, C-terminal / Factor H binding protein, C-terminal / Porin - #90 / Outer membrane protein/outer membrane enzyme PagP, beta-barrel /  Porin / cell outer membrane / Porin / cell outer membrane /  Beta Barrel / Mainly Beta / Factor H-binding protein Beta Barrel / Mainly Beta / Factor H-binding protein Function and homology information Function and homology information | ||||||
| Biological species |   NEISSERIA MENINGITIDIS MC58 (bacteria) NEISSERIA MENINGITIDIS MC58 (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.06 Å MOLECULAR REPLACEMENT / Resolution: 2.06 Å | ||||||
 Authors Authors | Johnson, S. / Tan, L. / van der Veen, S. / Caesar, J. / Goicoechea De Jorge, E. / Everett, R.J. / Bai, X. / Exley, R.M. / Ward, P.N. / Ruivo, N. ...Johnson, S. / Tan, L. / van der Veen, S. / Caesar, J. / Goicoechea De Jorge, E. / Everett, R.J. / Bai, X. / Exley, R.M. / Ward, P.N. / Ruivo, N. / Trivedi, K. / Cumber, E. / Jones, R. / Newham, L. / Staunton, D. / Borrow, R. / Pickering, M. / Lea, S.M. / Tang, C.M. | ||||||
 Citation Citation |  Journal: Plos Pathog. / Year: 2012 Journal: Plos Pathog. / Year: 2012Title: Design and Evaluation of Meningococcal Vaccines Through Structure-Based Modification of Host and Pathogen Molecules. Authors: Johnson, S. / Tan, L. / Van Der Veen, S. / Caesar, J. / Goicoechea De Jorge, E. / Harding, R.J. / Bai, X. / Exley, R.M. / Ward, P.N. / Ruivo, N. / Trivedi, K. / Cumber, E. / Jones, R. / ...Authors: Johnson, S. / Tan, L. / Van Der Veen, S. / Caesar, J. / Goicoechea De Jorge, E. / Harding, R.J. / Bai, X. / Exley, R.M. / Ward, P.N. / Ruivo, N. / Trivedi, K. / Cumber, E. / Jones, R. / Newham, L. / Staunton, D. / Ufret-Vincenty, R. / Borrow, R. / Pickering, M.C. / Lea, S.M. / Tang, C.M. | ||||||
| History |
| ||||||
| Remark 700 | SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS "AA" IN EACH CHAIN ON SHEET RECORDS BELOW ... SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS "AA" IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 8-STRANDED BARREL THIS IS REPRESENTED BY A 9-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. THE SHEETS PRESENTED AS "BA" IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 8-STRANDED BARREL THIS IS REPRESENTED BY A 9-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4ayn.cif.gz 4ayn.cif.gz | 69.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4ayn.ent.gz pdb4ayn.ent.gz | 48.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4ayn.json.gz 4ayn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ay/4ayn https://data.pdbj.org/pub/pdb/validation_reports/ay/4ayn ftp://data.pdbj.org/pub/pdb/validation_reports/ay/4ayn ftp://data.pdbj.org/pub/pdb/validation_reports/ay/4ayn | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2ybyC  4aydC  4ayeC  4ayiC  4aymC  2w81S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (0.31355, -0.94855, -0.04401), Vector  : : |
- Components
Components
| #1: Protein | Mass: 29540.922 Da / Num. of mol.: 2 / Fragment: RESIDUES 27-273 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   NEISSERIA MENINGITIDIS MC58 (bacteria) / Variant: P22 / Production host: NEISSERIA MENINGITIDIS MC58 (bacteria) / Variant: P22 / Production host:   ESCHERICHIA COLI (E. coli) / Strain (production host): B834(DE3) / References: UniProt: C6KHT4 ESCHERICHIA COLI (E. coli) / Strain (production host): B834(DE3) / References: UniProt: C6KHT4#2: Chemical | ChemComp-SO4 /  Sulfate Sulfate#3: Water | ChemComp-HOH / |  Water WaterSequence details | DISCREPANCIES AT TERMINI ARE FROM VECTOR. THE SEQUENCE HAS BEEN RENUMBERED TO MATCH THAT OF THE ...DISCREPANC | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.05 Å3/Da / Density % sol: 40 % / Description: NONE |
|---|---|
Crystal grow | pH: 4.6 Details: 30% PEG2KMME, 0.1 M SODIUM ACETATE PH 4.6, 0.2 M AMMONIUM SULPHATE |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID29 / Wavelength: 0.96 / Beamline: ID29 / Wavelength: 0.96 |
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Feb 28, 2011 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.96 Å / Relative weight: 1 : 0.96 Å / Relative weight: 1 |
| Reflection | Resolution: 2.06→60 Å / Num. obs: 14737 / % possible obs: 98.5 % / Observed criterion σ(I): 2 / Redundancy: 5.9 % / Biso Wilson estimate: 32.19 Å2 / Rmerge(I) obs: 0.04 / Net I/σ(I): 26 |
| Reflection shell | Resolution: 2.06→2.18 Å / Redundancy: 4.1 % / Rmerge(I) obs: 0.19 / Mean I/σ(I) obs: 6.8 / % possible all: 91.3 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2W81 Resolution: 2.06→15 Å / Cor.coef. Fo:Fc: 0.9491 / Cor.coef. Fo:Fc free: 0.9296 / SU R Cruickshank DPI: 0.208 / Cross valid method: THROUGHOUT / σ(F): 0 / SU R Blow DPI: 0.224 / SU Rfree Blow DPI: 0.168 / SU Rfree Cruickshank DPI: 0.164 Details: IDEAL-DIST CONTACT TERM CONTACT SETUP ALL ATOMS HAVE CCP4 ATOM TYPE FROM LIBRARY
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 32.68 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.22 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.06→15 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.06→2.23 Å / Total num. of bins used: 7
|
 Movie
Movie Controller
Controller











 PDBj
PDBj





