[English] 日本語
 Yorodumi
Yorodumi- PDB-3slm: Crystal structure of the 2'- Deoxyguanosine riboswitch bound to 2... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3slm | ||||||
|---|---|---|---|---|---|---|---|
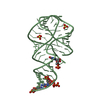
| Title | Crystal structure of the 2'- Deoxyguanosine riboswitch bound to 2'-deoxyguanosine-5'-monophosphate | ||||||
 Components Components | RNA (68-MER) | ||||||
 Keywords Keywords |  RNA / RNA /  three-way junction / three-way junction /  riboswitch / 2'-deoxy guanosine riboswitch / 2'-deoxy guanosine | ||||||
| Function / homology | 2'-DEOXYGUANOSINE-5'-MONOPHOSPHATE /  SUCCINIC ACID / SUCCINIC ACID /  RNA / RNA (> 10) RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Molrep / Resolution: 2.7 Å SYNCHROTRON / Molrep / Resolution: 2.7 Å | ||||||
 Authors Authors | Pikovskaya, O. / Polonskaia, A. / Patel, D.J. / Serganov, A.A. | ||||||
 Citation Citation |  Journal: Nat.Chem.Biol. / Year: 2011 Journal: Nat.Chem.Biol. / Year: 2011Title: Structural principles of nucleoside selectivity in a 2'-deoxyguanosine riboswitch. Authors: Pikovskaya, O. / Polonskaia, A. / Patel, D.J. / Serganov, A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3slm.cif.gz 3slm.cif.gz | 86.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3slm.ent.gz pdb3slm.ent.gz | 65.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3slm.json.gz 3slm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/sl/3slm https://data.pdbj.org/pub/pdb/validation_reports/sl/3slm ftp://data.pdbj.org/pub/pdb/validation_reports/sl/3slm ftp://data.pdbj.org/pub/pdb/validation_reports/sl/3slm | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3skiC  3sklC  3skrC  3sktC  3skwC  3skzC  3slqC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
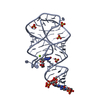
| 1 | 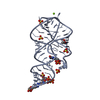
| ||||||||
| 2 | 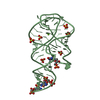
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
-RNA chain , 1 types, 2 molecules AB
| #1: RNA chain | Mass: 22018.873 Da / Num. of mol.: 2 / Source method: obtained synthetically / Details: in vitro transcribed |
|---|
-Non-polymers , 5 types, 29 molecules 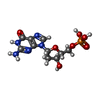








| #2: Chemical |  Deoxyguanosine monophosphate Deoxyguanosine monophosphate#3: Chemical | ChemComp-SO4 /  Sulfate Sulfate#4: Chemical | ChemComp-SIN / |  Succinic acid Succinic acid#5: Chemical | #6: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.3 Å3/Da / Density % sol: 46.63 % |
|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 5.1 Details: 0.05 M Na-succinate, ~3.0 M ammonium sulfate, 0.5 mM spermine and 20 mM magnesium chloride , pH 5.1, VAPOR DIFFUSION, HANGING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X29A / Wavelength: 1.075 Å / Beamline: X29A / Wavelength: 1.075 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Apr 27, 2010 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.075 Å / Relative weight: 1 : 1.075 Å / Relative weight: 1 |
| Reflection | Resolution: 2.7→20 Å / Num. all: 11084 / Num. obs: 11084 / % possible obs: 100 % / Redundancy: 4 % / Rmerge(I) obs: 0.096 / Net I/σ(I): 19.3 |
| Reflection shell | Resolution: 2.7→2.8 Å / Redundancy: 4.1 % / Rmerge(I) obs: 0.599 / Mean I/σ(I) obs: 2.6 / Num. unique all: 1100 / % possible all: 100 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : Molrep / Resolution: 2.7→19.543 Å / SU ML: 0.77 / σ(F): 1.34 / Phase error: 28.08 / Stereochemistry target values: ML : Molrep / Resolution: 2.7→19.543 Å / SU ML: 0.77 / σ(F): 1.34 / Phase error: 28.08 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.72 Å / VDW probe radii: 1 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 16.757 Å2 / ksol: 0.283 e/Å3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.7→19.543 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller








 PDBj
PDBj

































