[English] 日本語
 Yorodumi
Yorodumi- PDB-3q90: Crystal structure of the NTF2 domain of Ras GTPase-activating pro... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3q90 | ||||||
|---|---|---|---|---|---|---|---|
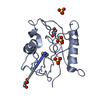
| Title | Crystal structure of the NTF2 domain of Ras GTPase-activating protein-binding protein 1 | ||||||
 Components Components | Ras GTPase-activating protein-binding protein 1 | ||||||
 Keywords Keywords |  HYDROLASE / HYDROLASE /  Structural Genomics / Structural Genomics /  Structural Genomics Consortium / SGC / NTF2-like (a+b proteins) / Protein Binding and Helicase / Protein (Ras GTPase-activating protein) / DNA and RNA binding / Structural Genomics Consortium / SGC / NTF2-like (a+b proteins) / Protein Binding and Helicase / Protein (Ras GTPase-activating protein) / DNA and RNA binding /  plasma membrane / plasma membrane /  nucleus nucleus | ||||||
| Function / homology |  Function and homology information Function and homology informationDNA/RNA helicase activity / positive regulation of stress granule assembly /  ribosomal small subunit binding / positive regulation of type I interferon production / ribosomal small subunit binding / positive regulation of type I interferon production /  stress granule assembly / stress granule assembly /  DNA helicase activity / molecular condensate scaffold activity / negative regulation of canonical Wnt signaling pathway / cytoplasmic stress granule / DNA helicase activity / molecular condensate scaffold activity / negative regulation of canonical Wnt signaling pathway / cytoplasmic stress granule /  perikaryon ...DNA/RNA helicase activity / positive regulation of stress granule assembly / perikaryon ...DNA/RNA helicase activity / positive regulation of stress granule assembly /  ribosomal small subunit binding / positive regulation of type I interferon production / ribosomal small subunit binding / positive regulation of type I interferon production /  stress granule assembly / stress granule assembly /  DNA helicase activity / molecular condensate scaffold activity / negative regulation of canonical Wnt signaling pathway / cytoplasmic stress granule / DNA helicase activity / molecular condensate scaffold activity / negative regulation of canonical Wnt signaling pathway / cytoplasmic stress granule /  perikaryon / perikaryon /  endonuclease activity / defense response to virus / endonuclease activity / defense response to virus /  DNA helicase / Ras protein signal transduction / DNA helicase / Ras protein signal transduction /  RNA helicase activity / RNA helicase activity /  RNA helicase / RNA helicase /  ribonucleoprotein complex / ribonucleoprotein complex /  focal adhesion / focal adhesion /  innate immune response / innate immune response /  mRNA binding / SARS-CoV-2 activates/modulates innate and adaptive immune responses / mRNA binding / SARS-CoV-2 activates/modulates innate and adaptive immune responses /  ATP hydrolysis activity / ATP hydrolysis activity /  DNA binding / DNA binding /  RNA binding / RNA binding /  ATP binding / ATP binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | ||||||
 Authors Authors | Welin, M. / Tresaugues, L. / Arrowsmith, C.H. / Berglund, H. / Bountra, C. / Collins, R. / Edwards, A.M. / Ekblad, T. / Flodin, S. / Flores, A. ...Welin, M. / Tresaugues, L. / Arrowsmith, C.H. / Berglund, H. / Bountra, C. / Collins, R. / Edwards, A.M. / Ekblad, T. / Flodin, S. / Flores, A. / Graslund, S. / Hammarstrom, M. / Johansson, I. / Karlberg, T. / Kol, S. / Kotenyova, T. / Kouznetsova, E. / Moche, M. / Nyman, T. / Persson, C. / Schuler, H. / Schutz, P. / Siponen, M.I. / Thorsell, A.G. / Van Der Berg, S. / Wahlberg, E. / Weigelt, J. / Nordlund, P. / Structural Genomics Consortium (SGC) | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Crystal structure of the NTF2 domain of Ras GTPase-activating protein-binding protein 1 Authors: Welin, M. / Tresaugues, L. / Arrowsmith, C.H. / Berglund, H. / Bountra, C. / Collins, R. / Edwards, A.M. / Ekblad, T. / Flodin, S. / Flores, A. / Graslund, S. / Hammarstrom, M. / Johansson, ...Authors: Welin, M. / Tresaugues, L. / Arrowsmith, C.H. / Berglund, H. / Bountra, C. / Collins, R. / Edwards, A.M. / Ekblad, T. / Flodin, S. / Flores, A. / Graslund, S. / Hammarstrom, M. / Johansson, I. / Karlberg, T. / Kol, S. / Kotenyova, T. / Kouznetsova, E. / Moche, M. / Nyman, T. / Persson, C. / Schuler, H. / Schutz, P. / Siponen, M.I. / Thorsell, A.G. / Van Der Berg, S. / Wahlberg, E. / Weigelt, J. / Nordlund, P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3q90.cif.gz 3q90.cif.gz | 115.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3q90.ent.gz pdb3q90.ent.gz | 91.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3q90.json.gz 3q90.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/q9/3q90 https://data.pdbj.org/pub/pdb/validation_reports/q9/3q90 ftp://data.pdbj.org/pub/pdb/validation_reports/q9/3q90 ftp://data.pdbj.org/pub/pdb/validation_reports/q9/3q90 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1z02S S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 15986.153 Da / Num. of mol.: 2 / Fragment: unp residues 1-139 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: G3BP, G3BP1 / Plasmid: pNIC-Bsa4 / Production host: Homo sapiens (human) / Gene: G3BP, G3BP1 / Plasmid: pNIC-Bsa4 / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3) R3 pRARE / References: UniProt: Q13283, Escherichia coli (E. coli) / Strain (production host): BL21(DE3) R3 pRARE / References: UniProt: Q13283,  DNA helicase, DNA helicase,  RNA helicase RNA helicase#2: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.12 Å3/Da / Density % sol: 41.97 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, sitting drop / pH: 5.5 Details: 25% w/v PEG3350, 0.2M AMMONIUM ACETATE, 0.1M BIS-TRIS, VAPOR DIFFUSION, SITTING DROP, temperature 277K, pH 5.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID23-2 / Wavelength: 0.8726 Å / Beamline: ID23-2 / Wavelength: 0.8726 Å |
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: Jun 20, 2010 / Details: MIRRORS |
| Radiation | Monochromator: horizontally side diffracting Silicon 111 crystal Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.8726 Å / Relative weight: 1 : 0.8726 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→50 Å / Num. all: 30636 / Num. obs: 30582 / % possible obs: 99.8 % / Observed criterion σ(I): -3 / Redundancy: 6.17 % / Biso Wilson estimate: 22.39 Å2 / Rsym value: 0.063 / Net I/σ(I): 17.17 |
| Reflection shell | Resolution: 1.7→1.8 Å / Redundancy: 3.67 % / Mean I/σ(I) obs: 3.48 / Num. unique all: 4585 / Rsym value: 0.434 / % possible all: 99.8 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1Z02 Resolution: 1.7→44.29 Å / Cor.coef. Fo:Fc: 0.9302 / Cor.coef. Fo:Fc free: 0.9175 / Cross valid method: THROUGHOUT / σ(F): 0
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 29.55 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.239 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→44.29 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.7→1.76 Å / Total num. of bins used: 15
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj

