+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3gva | ||||||
|---|---|---|---|---|---|---|---|
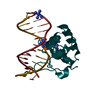
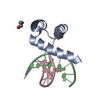
| Title | Crystal Structure Analysis of S. Pombe ATL | ||||||
 Components Components | Alkyltransferase-like protein 1 | ||||||
 Keywords Keywords |  DNA BINDING PROTEIN / Alkylated DNA damage repair / DNA BINDING PROTEIN / Alkylated DNA damage repair /  DNA damage / DNA damage /  DNA repair / DNA-binding DNA repair / DNA-binding | ||||||
| Function / homology |  Function and homology information Function and homology informationO6-alkylguanine-DNA binding / global genome nucleotide-excision repair / transcription-coupled nucleotide-excision repair /  catalytic activity / damaged DNA binding / catalytic activity / damaged DNA binding /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Schizosaccharomyces pombe (fission yeast) Schizosaccharomyces pombe (fission yeast) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 2 Å SYNCHROTRON / Resolution: 2 Å | ||||||
 Authors Authors | Tubbs, J.L. / Arvai, A.S. / Tainer, J.A. | ||||||
 Citation Citation |  Journal: Nature / Year: 2009 Journal: Nature / Year: 2009Title: Flipping of alkylated DNA damage bridges base and nucleotide excision repair. Authors: Tubbs, J.L. / Latypov, V. / Kanugula, S. / Butt, A. / Melikishvili, M. / Kraehenbuehl, R. / Fleck, O. / Marriott, A. / Watson, A.J. / Verbeek, B. / McGown, G. / Thorncroft, M. / Santibanez- ...Authors: Tubbs, J.L. / Latypov, V. / Kanugula, S. / Butt, A. / Melikishvili, M. / Kraehenbuehl, R. / Fleck, O. / Marriott, A. / Watson, A.J. / Verbeek, B. / McGown, G. / Thorncroft, M. / Santibanez-Koref, M.F. / Millington, C. / Arvai, A.S. / Kroeger, M.D. / Peterson, L.A. / Williams, D.M. / Fried, M.G. / Margison, G.P. / Pegg, A.E. / Tainer, J.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3gva.cif.gz 3gva.cif.gz | 57.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3gva.ent.gz pdb3gva.ent.gz | 42.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3gva.json.gz 3gva.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/gv/3gva https://data.pdbj.org/pub/pdb/validation_reports/gv/3gva ftp://data.pdbj.org/pub/pdb/validation_reports/gv/3gva ftp://data.pdbj.org/pub/pdb/validation_reports/gv/3gva | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 13662.412 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Schizosaccharomyces pombe (fission yeast) Schizosaccharomyces pombe (fission yeast)Gene: atl1, SPAC1250.04c / Production host:   Escherichia coli (E. coli) / References: UniProt: Q9UTN9 Escherichia coli (E. coli) / References: UniProt: Q9UTN9#2: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.92 Å3/Da / Density % sol: 35.83 % |
|---|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ALS ALS  / Beamline: 12.3.1 / Beamline: 12.3.1 |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Sep 15, 2007 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
| Reflection | Resolution: 2→50 Å / Num. obs: 14855 / % possible obs: 98.7 % / Redundancy: 9.3 % / Biso Wilson estimate: 14.7 Å2 / Rmerge(I) obs: 0.071 / Rsym value: 0.071 / Net I/σ(I): 25.2 |
| Reflection shell | Resolution: 2→2.7 Å / Redundancy: 8.8 % / Rmerge(I) obs: 0.286 / Mean I/σ(I) obs: 4.9 / Rsym value: 0.286 / % possible all: 98.5 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2→39.02 Å / Rfactor Rfree error: 0.007 / Occupancy max: 1 / Occupancy min: 1 / Data cutoff high absF: 114432 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Details: BULK SOLVENT MODEL USED
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 41.741 Å2 / ksol: 0.35 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 98.77 Å2 / Biso mean: 34.943 Å2 / Biso min: 13.99 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→39.02 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.13 Å / Rfactor Rfree error: 0.022 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj
