Entry Database : PDB / ID : 2vzvTitle Substrate Complex of Amycolatopsis orientalis exo-chitosanase CsxA E541A with chitosan EXO-BETA-D-GLUCOSAMINIDASE Keywords / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species AMYCOLATOPSIS ORIENTALIS (bacteria)Method / / / Resolution : 2.7 Å Authors Lammerts van Bueren, A. / Ghinet, M.G. / Gregg, K. / Fleury, A. / Brzezinski, R. / Boraston, A.B. Journal : J.Mol.Biol. / Year : 2009Title : The Structural Basis of Substrate Recognition in an Exo-Beta-D-Glucosaminidase Involved in Chitosan Hydrolysis.Authors : Lammerts Van Bueren, A. / Ghinet, M.G. / Gregg, K. / Fleury, A. / Brzezinski, R. / Boraston, A.B. History Deposition Aug 5, 2008 Deposition site / Processing site Revision 1.0 Oct 14, 2008 Provider / Type Revision 1.1 Aug 17, 2011 Group Atomic model / Derived calculations ... Atomic model / Derived calculations / Non-polymer description / Other / Refinement description / Structure summary / Version format compliance Revision 2.0 Jul 29, 2020 Group Advisory / Atomic model ... Advisory / Atomic model / Data collection / Derived calculations / Other / Structure summary Category atom_site / chem_comp ... atom_site / chem_comp / entity / pdbx_branch_scheme / pdbx_chem_comp_identifier / pdbx_database_status / pdbx_entity_branch / pdbx_entity_branch_descriptor / pdbx_entity_branch_link / pdbx_entity_branch_list / pdbx_entity_nonpoly / pdbx_nonpoly_scheme / pdbx_struct_assembly_gen / pdbx_unobs_or_zero_occ_atoms / struct_asym / struct_conn / struct_site / struct_site_gen Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_asym_id / _atom_site.auth_atom_id / _atom_site.auth_seq_id / _atom_site.label_asym_id / _atom_site.label_atom_id / _atom_site.type_symbol / _chem_comp.name / _chem_comp.type / _entity.formula_weight / _entity.pdbx_description / _entity.pdbx_number_of_molecules / _entity.type / _pdbx_database_status.status_code_sf / _pdbx_struct_assembly_gen.asym_id_list / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id Description / Provider / Type
Show all Show less Remark 700 SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN ... SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN THE SHEET RECORDS BELOW, TWO SHEETS ARE DEFINED.
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
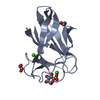
Components Exo-1,4-beta-D-glucosaminidase
Exo-1,4-beta-D-glucosaminidase  Keywords
Keywords HYDROLASE /
HYDROLASE /  EXO-BETA-D-GLUCOSAMINIDASE / GH2 /
EXO-BETA-D-GLUCOSAMINIDASE / GH2 /  CSXA /
CSXA /  CHITOSAN /
CHITOSAN /  GLYCOSIDE HYDROLASE
GLYCOSIDE HYDROLASE Function and homology information
Function and homology information exo-1,4-beta-D-glucosaminidase /
exo-1,4-beta-D-glucosaminidase /  exo-1,4-beta-D-glucosaminidase activity / chitin catabolic process / polysaccharide catabolic process / hydrolase activity, hydrolyzing O-glycosyl compounds /
exo-1,4-beta-D-glucosaminidase activity / chitin catabolic process / polysaccharide catabolic process / hydrolase activity, hydrolyzing O-glycosyl compounds /  carbohydrate binding / extracellular region
carbohydrate binding / extracellular region
 AMYCOLATOPSIS ORIENTALIS (bacteria)
AMYCOLATOPSIS ORIENTALIS (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON / OTHER / Resolution: 2.7 Å
SYNCHROTRON / OTHER / Resolution: 2.7 Å  Authors
Authors Citation
Citation Journal: J.Mol.Biol. / Year: 2009
Journal: J.Mol.Biol. / Year: 2009 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2vzv.cif.gz
2vzv.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2vzv.ent.gz
pdb2vzv.ent.gz PDB format
PDB format 2vzv.json.gz
2vzv.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/vz/2vzv
https://data.pdbj.org/pub/pdb/validation_reports/vz/2vzv ftp://data.pdbj.org/pub/pdb/validation_reports/vz/2vzv
ftp://data.pdbj.org/pub/pdb/validation_reports/vz/2vzv Links
Links Assembly
Assembly


 Components
Components Exo-1,4-beta-D-glucosaminidase / CSXA
Exo-1,4-beta-D-glucosaminidase / CSXA
 AMYCOLATOPSIS ORIENTALIS (bacteria) / Plasmid: PFD666 / Production host:
AMYCOLATOPSIS ORIENTALIS (bacteria) / Plasmid: PFD666 / Production host: 
 STREPTOMYCES LIVIDANS (bacteria) / Strain (production host): TK24 / References: UniProt: Q56F26
STREPTOMYCES LIVIDANS (bacteria) / Strain (production host): TK24 / References: UniProt: Q56F26 / Mass: 340.327 Da / Num. of mol.: 2
/ Mass: 340.327 Da / Num. of mol.: 2 Water
Water X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  NSLS
NSLS  / Beamline: X8C / Wavelength: 1.1
/ Beamline: X8C / Wavelength: 1.1  : 1.1 Å / Relative weight: 1
: 1.1 Å / Relative weight: 1  Processing
Processing : OTHER / Resolution: 2.7→185.7 Å / Cor.coef. Fo:Fc: 0.923 / Cor.coef. Fo:Fc free: 0.853 / SU B: 12.465 / SU ML: 0.257 / Cross valid method: THROUGHOUT / ESU R Free: 0.387 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
: OTHER / Resolution: 2.7→185.7 Å / Cor.coef. Fo:Fc: 0.923 / Cor.coef. Fo:Fc free: 0.853 / SU B: 12.465 / SU ML: 0.257 / Cross valid method: THROUGHOUT / ESU R Free: 0.387 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS. Movie
Movie Controller
Controller


















 PDBj
PDBj

