[English] 日本語
 Yorodumi
Yorodumi- PDB-2rcn: Crystal Structure of the Ribosomal interacting GTPase YjeQ from t... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2rcn | ||||||
|---|---|---|---|---|---|---|---|
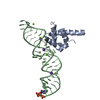
| Title | Crystal Structure of the Ribosomal interacting GTPase YjeQ from the Enterobacterial species Salmonella Typhimurium. | ||||||
 Components Components | Probable GTPase engC | ||||||
 Keywords Keywords |  HYDROLASE / YjeQ / HYDROLASE / YjeQ /  gtpase / circularly permuted / GTP-binding / Nucleotide-binding gtpase / circularly permuted / GTP-binding / Nucleotide-binding | ||||||
| Function / homology |  Function and homology information Function and homology information Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides / Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides /  ribosomal small subunit biogenesis / ribosomal small subunit biogenesis /  rRNA binding / rRNA binding /  GTPase activity / GTP binding / GTPase activity / GTP binding /  metal ion binding / metal ion binding /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Salmonella typhimurium (bacteria) Salmonella typhimurium (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.25 Å MOLECULAR REPLACEMENT / Resolution: 2.25 Å | ||||||
 Authors Authors | Nichols, C.E. / Stammers, D.K. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.F / Year: 2007 Journal: Acta Crystallogr.,Sect.F / Year: 2007Title: Structure of the ribosomal interacting GTPase YjeQ from the enterobacterial species Salmonella typhimurium. Authors: Nichols, C.E. / Johnson, C. / Lamb, H.K. / Lockyer, M. / Charles, I.G. / Hawkins, A.R. / Stammers, D.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2rcn.cif.gz 2rcn.cif.gz | 79 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2rcn.ent.gz pdb2rcn.ent.gz | 56.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2rcn.json.gz 2rcn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/rc/2rcn https://data.pdbj.org/pub/pdb/validation_reports/rc/2rcn ftp://data.pdbj.org/pub/pdb/validation_reports/rc/2rcn ftp://data.pdbj.org/pub/pdb/validation_reports/rc/2rcn | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| 2 | 
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
| |||||||||
| Details | The crystallographic monomer is equivalent to the biological monomer. |
- Components
Components
| #1: Protein | Mass: 39895.836 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Salmonella typhimurium (bacteria) / Strain: LT2 / Gene: engC / Plasmid: pMUT99 / Species (production host): Escherichia coli / Production host: Salmonella typhimurium (bacteria) / Strain: LT2 / Gene: engC / Plasmid: pMUT99 / Species (production host): Escherichia coli / Production host:   Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21DE3 Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21DE3References: UniProt: Q8ZKB0,  Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides |
|---|---|
| #2: Chemical | ChemComp-ZN / |
| #3: Chemical | ChemComp-MG / |
| #4: Chemical | ChemComp-GDP /  Guanosine diphosphate Guanosine diphosphate |
| #5: Water | ChemComp-HOH /  Water Water |
| Nonpolymer details | LIGAND GDP HAS WRONG STEREOCHEM |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.19 Å3/Da / Density % sol: 43.83 % |
|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 6 Details: 1.85 M ammonium sulphate, 0.1 M MES and 10 mM cobalt(II)chloride, pH 6.0, VAPOR DIFFUSION, SITTING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU FR-E+ DW / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU FR-E+ DW / Wavelength: 1.5418 Å |
| Detector | Type: MAR scanner 345 mm plate / Detector: IMAGE PLATE / Date: Jan 5, 2005 / Details: Osmic multilayer optics |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.25→30 Å / Num. all: 16948 / Num. obs: 16829 / % possible obs: 99.3 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -2 / Redundancy: 6.1 % / Biso Wilson estimate: 32.6 Å2 / Rmerge(I) obs: 0.104 / Net I/σ(I): 16.75 |
| Reflection shell | Resolution: 2.25→2.33 Å / Redundancy: 3.5 % / Rmerge(I) obs: 0.52 / Mean I/σ(I) obs: 1.99 / Num. unique all: 1672 / % possible all: 94.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1U0L and 1T9H Resolution: 2.25→27.89 Å / Rfactor Rfree error: 0.007 / Data cutoff high absF: 2438805.43 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): -2 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 61.7796 Å2 / ksol: 0.359131 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 39.9 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.25→27.89 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.25→2.33 Å / Rfactor Rfree error: 0.026 / Total num. of bins used: 10
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj






