Entry Database : PDB / ID : 2miqTitle Solution NMR Structure of PHD Type 1 Zinc Finger Domain 1 of Lysine-specific Demethylase Lid from Drosophila melanogaster, Northeast Structural Genomics Consortium (NESG) Target FR824J Lysine-specific demethylase lid Keywords / / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Drosophila melanogaster (fruit fly)Method / Model details lowest energy, model1 Authors Xu, X. / Eletsky, A. / Shastry, R. / Maglaqui, M. / Janjua, H. / Xiao, R. / Everett, J.K. / Sukumaran, D.K. / Montelione, G.T. / Szyperski, T. ...Xu, X. / Eletsky, A. / Shastry, R. / Maglaqui, M. / Janjua, H. / Xiao, R. / Everett, J.K. / Sukumaran, D.K. / Montelione, G.T. / Szyperski, T. / Northeast Structural Genomics Consortium (NESG) / Chaperone-Enabled Studies of Epigenetic Regulation Enzymes (CEBS) Journal : To be Published Title : Solution NMR Structure of PHD Type 1 Zinc Finger Domain 1 from Lysine-specific Demethylase Lid from Drosophila melanogaster, Northeast Structural Genomics Consortium (NESG) Target FR824JAuthors : Xu, X. / Eletsky, A. / Shastry, R. / Maglaqui, M. / Janjua, H. / Xiao, R. / Everett, J.K. / Sukumaran, D. / Montelione, G.T. / Szyperski, T. History Deposition Dec 17, 2013 Deposition site / Processing site Revision 1.0 Jan 29, 2014 Provider / Type Revision 1.1 Feb 19, 2014 Group Revision 1.2 Apr 23, 2014 Group / Structure summaryRevision 1.3 Jun 14, 2023 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Other Category database_2 / pdbx_database_status ... database_2 / pdbx_database_status / pdbx_nmr_software / pdbx_struct_conn_angle / struct_conn / struct_ref_seq_dif / struct_site Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_nmr_data / _pdbx_nmr_software.name / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_ref_seq_dif.details / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords OXIDOREDUCTASE /
OXIDOREDUCTASE /  Zinc finger / PSI-Biology /
Zinc finger / PSI-Biology /  Structural Genomics / Northeast Structural Genomics Consortium / NESG / Chaperone-Enabled Studies of Epigenetic Regulation Enzymes /
Structural Genomics / Northeast Structural Genomics Consortium / NESG / Chaperone-Enabled Studies of Epigenetic Regulation Enzymes /  CEBS
CEBS Function and homology information
Function and homology information enzyme inhibitor activity ...TFAP2 (AP-2) family regulates transcription of cell cycle factors / histone H4R3 demethylase activity / male germ-line stem cell population maintenance / synaptonemal complex organization / oocyte karyosome formation / larval somatic muscle development / [histone H3]-trimethyl-L-lysine4 demethylase / histone H3K4me/H3K4me2/H3K4me3 demethylase activity / negative regulation of stem cell differentiation /
enzyme inhibitor activity ...TFAP2 (AP-2) family regulates transcription of cell cycle factors / histone H4R3 demethylase activity / male germ-line stem cell population maintenance / synaptonemal complex organization / oocyte karyosome formation / larval somatic muscle development / [histone H3]-trimethyl-L-lysine4 demethylase / histone H3K4me/H3K4me2/H3K4me3 demethylase activity / negative regulation of stem cell differentiation /  enzyme inhibitor activity / Sin3-type complex / locomotor rhythm / histone reader activity / circadian regulation of gene expression / chromatin organization /
enzyme inhibitor activity / Sin3-type complex / locomotor rhythm / histone reader activity / circadian regulation of gene expression / chromatin organization /  chromatin remodeling /
chromatin remodeling /  chromatin / regulation of DNA-templated transcription / positive regulation of DNA-templated transcription / positive regulation of transcription by RNA polymerase II /
chromatin / regulation of DNA-templated transcription / positive regulation of DNA-templated transcription / positive regulation of transcription by RNA polymerase II /  DNA binding /
DNA binding /  metal ion binding /
metal ion binding /  nucleus
nucleus
 Drosophila melanogaster (fruit fly)
Drosophila melanogaster (fruit fly) SOLUTION NMR /
SOLUTION NMR /  simulated annealing
simulated annealing  Authors
Authors Citation
Citation Journal: To be Published
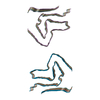
Journal: To be Published Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2miq.cif.gz
2miq.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2miq.ent.gz
pdb2miq.ent.gz PDB format
PDB format 2miq.json.gz
2miq.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/mi/2miq
https://data.pdbj.org/pub/pdb/validation_reports/mi/2miq ftp://data.pdbj.org/pub/pdb/validation_reports/mi/2miq
ftp://data.pdbj.org/pub/pdb/validation_reports/mi/2miq Links
Links Assembly
Assembly
 Components
Components
 Drosophila melanogaster (fruit fly) / Gene: CG9088, lid / Production host:
Drosophila melanogaster (fruit fly) / Gene: CG9088, lid / Production host: 
 Escherichia coli (E. coli) / Strain (production host): BL21(DE3)pMgK
Escherichia coli (E. coli) / Strain (production host): BL21(DE3)pMgK Oxidoreductases; Acting on paired donors, with incorporation or reduction of molecular oxygen; With 2-oxoglutarate as one donor, and incorporation of one atom of oxygen into each donor
Oxidoreductases; Acting on paired donors, with incorporation or reduction of molecular oxygen; With 2-oxoglutarate as one donor, and incorporation of one atom of oxygen into each donor SOLUTION NMR
SOLUTION NMR Sample preparation
Sample preparation Processing
Processing Movie
Movie Controller
Controller







 PDBj
PDBj


