+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2lmu | ||||||
|---|---|---|---|---|---|---|---|
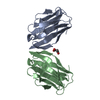
| Title | Androcam at high calcium | ||||||
 Components Components | Calmodulin-related protein 97A | ||||||
 Keywords Keywords | METAL BINDING PROTEIN /  spermatogenesis spermatogenesis | ||||||
| Function / homology |  Function and homology information Function and homology informationinvestment cone /  myosin VI complex / myosin VI head/neck binding / actin filament-based movement / Neutrophil degranulation / enzyme regulator activity / myosin VI complex / myosin VI head/neck binding / actin filament-based movement / Neutrophil degranulation / enzyme regulator activity /  calcium ion binding / calcium ion binding /  nucleus / nucleus /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
| Model details | lowest energy, model 1 | ||||||
 Authors Authors | Joshi, M.K. / Moran, S.T. / Beckingham, K.M. / Mackenzie, K.R. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2012 Journal: Proc.Natl.Acad.Sci.USA / Year: 2012Title: Structure of androcam supports specialized interactions with myosin VI. Authors: Joshi, M.K. / Moran, S. / Beckingham, K.M. / Mackenzie, K.R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2lmu.cif.gz 2lmu.cif.gz | 998.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2lmu.ent.gz pdb2lmu.ent.gz | 875 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2lmu.json.gz 2lmu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lm/2lmu https://data.pdbj.org/pub/pdb/validation_reports/lm/2lmu ftp://data.pdbj.org/pub/pdb/validation_reports/lm/2lmu ftp://data.pdbj.org/pub/pdb/validation_reports/lm/2lmu | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 17028.871 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Drosophila melanogaster (fruit fly) / Gene: And, Camr97A, CG17769 / Production host: Drosophila melanogaster (fruit fly) / Gene: And, Camr97A, CG17769 / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: P49258 Escherichia coli (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: P49258 |
|---|---|
| #2: Chemical |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR / Details: structure of androcam in high (10mM) calcium SOLUTION NMR / Details: structure of androcam in high (10mM) calcium | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 1.8 mM [U-100% 13C; U-100% 15N] androcam, 10 mM calcium chloride, 10 % D2O, 10 mM potassium chloride, 10 mM TRIS, 10 mM DTT, 0.1 mM TSP, 90% H2O/10% D2O Solvent system: 90% H2O/10% D2O | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||||||
| Sample conditions | Ionic strength: 10 / pH: 7.4 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 | ||||||||||||||||||||||||
| NMR constraints | Protein chi angle constraints total count: 55 / Protein phi angle constraints total count: 103 / Protein psi angle constraints total count: 103 | ||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||
| NMR ensemble | Average torsion angle constraint violation: 0.3 ° Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 20 / Maximum torsion angle constraint violation: 1 ° |
 Movie
Movie Controller
Controller












 PDBj
PDBj



