[English] 日本語
 Yorodumi
Yorodumi- PDB-2kz3: Backbone 1H, 13C, and 15N Chemical Shift Assignments for human Ra... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2kz3 | ||||||
|---|---|---|---|---|---|---|---|
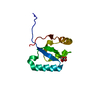
| Title | Backbone 1H, 13C, and 15N Chemical Shift Assignments for human Rad51D from 1 to 83 | ||||||
 Components Components | Putative uncharacterized protein RAD51L3 | ||||||
 Keywords Keywords | UNKNOWN FUNCTION / Rad51D /  Homologous Recombination Homologous Recombination | ||||||
| Function / homology |  Function and homology information Function and homology informationRad51B-Rad51C-Rad51D-XRCC2 complex / DNA strand invasion / telomere maintenance via recombination / Impaired BRCA2 binding to PALB2 / gamma-tubulin binding / reciprocal meiotic recombination / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Homologous DNA Pairing and Strand Exchange ...Rad51B-Rad51C-Rad51D-XRCC2 complex / DNA strand invasion / telomere maintenance via recombination / Impaired BRCA2 binding to PALB2 / gamma-tubulin binding / reciprocal meiotic recombination / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Homologous DNA Pairing and Strand Exchange / Resolution of D-loop Structures through Synthesis-Dependent Strand Annealing (SDSA) / Resolution of D-loop Structures through Holliday Junction Intermediates / ATP-dependent DNA damage sensor activity / Presynaptic phase of homologous DNA pairing and strand exchange / ATP-dependent activity, acting on DNA / interstrand cross-link repair /  telomere maintenance / telomere maintenance /  replication fork / TP53 Regulates Transcription of DNA Repair Genes / double-strand break repair via homologous recombination / HDR through Homologous Recombination (HRR) / replication fork / TP53 Regulates Transcription of DNA Repair Genes / double-strand break repair via homologous recombination / HDR through Homologous Recombination (HRR) /  single-stranded DNA binding / single-stranded DNA binding /  chromosome, telomeric region / chromosome, telomeric region /  regulation of cell cycle / regulation of cell cycle /  DNA repair / DNA repair /  centrosome / centrosome /  ATP hydrolysis activity / ATP hydrolysis activity /  DNA binding / DNA binding /  nucleoplasm / nucleoplasm /  ATP binding / ATP binding /  nucleus / nucleus /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / torsion angle dynamics SOLUTION NMR / torsion angle dynamics | ||||||
| Model details | lowest energy, model 1 | ||||||
 Authors Authors | Choi, N. / Kim, Y. | ||||||
 Citation Citation |  Journal: Int.J.Biochem.Cell Biol. / Year: 2011 Journal: Int.J.Biochem.Cell Biol. / Year: 2011Title: Structural and functional characterization of the N-terminal domain of human Rad51D Authors: Kim, Y.M. / Choi, B.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2kz3.cif.gz 2kz3.cif.gz | 508 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2kz3.ent.gz pdb2kz3.ent.gz | 425.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2kz3.json.gz 2kz3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kz/2kz3 https://data.pdbj.org/pub/pdb/validation_reports/kz/2kz3 ftp://data.pdbj.org/pub/pdb/validation_reports/kz/2kz3 ftp://data.pdbj.org/pub/pdb/validation_reports/kz/2kz3 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 9070.666 Da / Num. of mol.: 1 / Fragment: UNP residues 1-83 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) / References: UniProt: C9JSQ7, UniProt: O75771*PLUS Escherichia coli (E. coli) / References: UniProt: C9JSQ7, UniProt: O75771*PLUS |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sample conditions | Ionic strength: 100 / pH: 7.5 / Pressure: 1 atm / Temperature: 287 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software | Name:  CYANA / Version: 2.1 / Developer: P.GUNTERT ET AL. / Classification: refinement CYANA / Version: 2.1 / Developer: P.GUNTERT ET AL. / Classification: refinement |
|---|---|
| Refinement | Method: torsion angle dynamics / Software ordinal: 1 |
| NMR representative | Selection criteria: lowest energy |
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations Conformers calculated total number: 100 / Conformers submitted total number: 20 / Representative conformer: 1 |
 Movie
Movie Controller
Controller









 PDBj
PDBj







