+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2kjy | ||||||
|---|---|---|---|---|---|---|---|
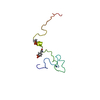
| Title | MYPT1(658-714) | ||||||
 Components Components | Protein phosphatase 1 regulatory subunit 12A | ||||||
 Keywords Keywords |  SIGNALING PROTEIN / MYPT1 / SIGNALING PROTEIN / MYPT1 /  phosphorylation / phosphorylation /  protein phosphatase 1 / myosin phosphatase / protein phosphatase 1 / myosin phosphatase /  Alternative splicing / ANK repeat / Alternative splicing / ANK repeat /  Cytoplasm / Cytoplasm /  Phosphoprotein / Polymorphism Phosphoprotein / Polymorphism | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of myosin-light-chain-phosphatase activity / positive regulation of myosin-light-chain-phosphatase activity / phosphatase regulator activity / PTW/PP1 phosphatase complex / contractile muscle fiber / regulation of nucleocytoplasmic transport / negative regulation of catalytic activity / RHO GTPases Activate ROCKs / RHO GTPases activate CIT /  enzyme inhibitor activity ...regulation of myosin-light-chain-phosphatase activity / positive regulation of myosin-light-chain-phosphatase activity / phosphatase regulator activity / PTW/PP1 phosphatase complex / contractile muscle fiber / regulation of nucleocytoplasmic transport / negative regulation of catalytic activity / RHO GTPases Activate ROCKs / RHO GTPases activate CIT / enzyme inhibitor activity ...regulation of myosin-light-chain-phosphatase activity / positive regulation of myosin-light-chain-phosphatase activity / phosphatase regulator activity / PTW/PP1 phosphatase complex / contractile muscle fiber / regulation of nucleocytoplasmic transport / negative regulation of catalytic activity / RHO GTPases Activate ROCKs / RHO GTPases activate CIT /  enzyme inhibitor activity / enzyme inhibitor activity /  centrosome cycle / A band / RHO GTPases activate PAKs / centrosome cycle / A band / RHO GTPases activate PAKs /  regulation of cell adhesion / regulation of cell adhesion /  stress fiber / RHO GTPases activate PKNs / 14-3-3 protein binding / protein dephosphorylation / stress fiber / RHO GTPases activate PKNs / 14-3-3 protein binding / protein dephosphorylation /  kinetochore / Z disc / kinetochore / Z disc /  Regulation of PLK1 Activity at G2/M Transition / Regulation of PLK1 Activity at G2/M Transition /  actin cytoskeleton / cellular response to xenobiotic stimulus / mitotic cell cycle / actin cytoskeleton / cellular response to xenobiotic stimulus / mitotic cell cycle /  focal adhesion / focal adhesion /  centrosome / centrosome /  nucleolus / nucleolus /  protein kinase binding / protein kinase binding /  signal transduction / positive regulation of transcription by RNA polymerase II / signal transduction / positive regulation of transcription by RNA polymerase II /  nucleoplasm / nucleoplasm /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / DGSA-distance geometry simulated annealing SOLUTION NMR / DGSA-distance geometry simulated annealing | ||||||
| Model details | minimized average, model 1 | ||||||
 Authors Authors | Mori, S. / Iwaoka, R. / Eto, M. / Ohki, S. | ||||||
 Citation Citation |  Journal: Proteins / Year: 2009 Journal: Proteins / Year: 2009Title: Solution structure of the inhibitory phosphorylation domain of myosin phosphatase targeting subunit 1 Authors: Mori, S. / Iwaoka, R. / Eto, M. / Ohki, S.Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2kjy.cif.gz 2kjy.cif.gz | 381.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2kjy.ent.gz pdb2kjy.ent.gz | 319.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2kjy.json.gz 2kjy.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kj/2kjy https://data.pdbj.org/pub/pdb/validation_reports/kj/2kjy ftp://data.pdbj.org/pub/pdb/validation_reports/kj/2kjy ftp://data.pdbj.org/pub/pdb/validation_reports/kj/2kjy | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 6964.675 Da / Num. of mol.: 1 / Fragment: UNP residues 658- 714 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: PPP1R12A, MBS, MYPT1 / Plasmid: pET / Production host: Homo sapiens (human) / Gene: PPP1R12A, MBS, MYPT1 / Plasmid: pET / Production host:   Escherichia coli (E. coli) / References: UniProt: O14974 Escherichia coli (E. coli) / References: UniProt: O14974 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 0.6-1.0mM MYPT1(658-714)-1, 0.6-1.0mM [U-99% 15N] MYPT1(658-714)-2, 0.6-1.0mM [U-99% 13C; U-99% 15N] MYPT1(658-714)-3, 90% H2O/10% D2O Solvent system: 90% H2O/10% D2O | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||
| Sample conditions | Ionic strength: KCl @100 / pH: 6.0 / Pressure: ambient / Temperature: 285 K |
-NMR measurement
| NMR spectrometer | Type: Varian INOVA / Manufacturer: Varian / Model : INOVA / Field strength: 750 MHz : INOVA / Field strength: 750 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: DGSA-distance geometry simulated annealing / Software ordinal: 1 | ||||||||||||
| NMR representative | Selection criteria: minimized average | ||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 100 / Conformers submitted total number: 20 / Representative conformer: 1 |
 Movie
Movie Controller
Controller








 PDBj
PDBj





