+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2k0y | ||||||
|---|---|---|---|---|---|---|---|
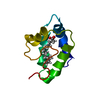
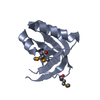
| Title | The actinorhodin apo acyl carrier protein from S. coelicolor | ||||||
 Components Components | Actinorhodin polyketide synthase acyl carrier protein | ||||||
 Keywords Keywords |  TRANSPORT PROTEIN / TRANSPORT PROTEIN /  Acyl carrier Protein / Acyl carrier Protein /  Actinorhodin / Actinorhodin /  Polyketide / Polyketide /  Antibiotic / Antibiotic biosynthesis / Antibiotic / Antibiotic biosynthesis /  Phosphopantetheine Phosphopantetheine | ||||||
| Function / homology |  Function and homology information Function and homology informationlipid A biosynthetic process /  acyl binding / antibiotic biosynthetic process / acyl carrier activity / acyl binding / antibiotic biosynthetic process / acyl carrier activity /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Streptomyces coelicolor (bacteria) Streptomyces coelicolor (bacteria) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
| Model details | This is an ensemble of 20 NMR structures, ARIA 1.2 restraint files (ambiguous and non-ambiguous ...This is an ensemble of 20 NMR structures, ARIA 1.2 restraint files (ambiguous and non-ambiguous NOEs, TALOS restraints and J-coupling restraints. Chemical shift data has also been deposited for 1H, 15N and 13C. | ||||||
 Authors Authors | Crump, M.P. / Evans, S.E. / Christopher, W. | ||||||
 Citation Citation |  Journal: Chembiochem / Year: 2008 Journal: Chembiochem / Year: 2008Title: An ACP Structural Switch: Conformational Differences between the Apo and Holo Forms of the Actinorhodin Polyketide Synthase Acyl Carrier Protein. Authors: Evans, S.E. / Williams, C. / Arthur, C.J. / Burston, S.G. / Simpson, T.J. / Crosby, J. / Crump, M.P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2k0y.cif.gz 2k0y.cif.gz | 543.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2k0y.ent.gz pdb2k0y.ent.gz | 464.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2k0y.json.gz 2k0y.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/k0/2k0y https://data.pdbj.org/pub/pdb/validation_reports/k0/2k0y ftp://data.pdbj.org/pub/pdb/validation_reports/k0/2k0y ftp://data.pdbj.org/pub/pdb/validation_reports/k0/2k0y | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 9239.177 Da / Num. of mol.: 1 / Mutation: C17S Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Streptomyces coelicolor (bacteria) Streptomyces coelicolor (bacteria)Description: Actinorhodin acyl carrier protein (act ACP) from S. coelicolor was heterologously overexpressed in its apo form in E. coli BL21 (DE3) cells. These cells contained the plasmid pET11c C17S ...Description: Actinorhodin acyl carrier protein (act ACP) from S. coelicolor was heterologously overexpressed in its apo form in E. coli BL21 (DE3) cells. These cells contained the plasmid pET11c C17S act ACP (courtesy of Dr. Tom Nicholson). This IPTG inducible vector is both easier to use and more reliable than the heat inducible pT7-7 version originally constructed. Production host:   Escherichia coli (E. coli) / Strain (production host): BL21 / Variant (production host): DE3 / References: UniProt: Q02054 Escherichia coli (E. coli) / Strain (production host): BL21 / Variant (production host): DE3 / References: UniProt: Q02054 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMRDetails: This is an ensemble of 20 NMR structures, ARIA 1.2 restraint files (ambiguous and non-ambiguous NOEs, TALOS restraints and J-coupling restraints. Chemical shift data has also been deposited for 1H, 15N and 13C. | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||
| NMR details | Text: The structure was determined using a combination of NOE data (ambiguous and unambiguous as determined by ARIA) combined with J-coupling and TALOS dihedral restraints |
- Sample preparation
Sample preparation
| Details | Contents: 1-2 mM [U-98% 13C; U-98% 15N] act ACP, 5% D2O, 95% H2O, 20 mM potassium phosphate, 1 mM sodium azide, 95% H2O/5% D2O Solvent system: 95% H2O/5% D2O | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||
| Sample conditions | pH: 5.5 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Varian INOVA / Manufacturer: Varian / Model : INOVA / Field strength: 600 MHz : INOVA / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 Details: All structure calculations were carried out using the Ambiguous Restraints for Iterative Assignment of NOEs (ARIA) protocol Version 1.2, which includes an algorithm that attempts to correct ...Details: All structure calculations were carried out using the Ambiguous Restraints for Iterative Assignment of NOEs (ARIA) protocol Version 1.2, which includes an algorithm that attempts to correct for the effects of spin diffusion, was use. Torsion Angle Likelihood Obtained from Shift and sequence similarity (TALOS) and was used to predict φ and ψ dihedral angle restraints. Initially, structure calculation runs contained 8 iterations of 20 structures each, with the best 7 structures in each iteration (sorted according to total energy) being used for analysis and assignment. The number of dynamics steps was increased over default values to 20000 and 16000 for the first and second cooling stages respectively. After each run, violated restraints were checked, and those arising from noise peaks or incorrect assignments were removed/reassigned. Final ensembles of 100 structures were calculated from calibrated restraint tables. The 20 best structures (sorted according to total energy) were selected for water refinement. Water refined structures were calculated using the slightly modified refinement script applied to the RECOORD database. PROCHECK and WHATCHECK and quality indicators were compared to the average values for the RECOORD database of protein NMR structures. | ||||||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 20 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller














 PDBj
PDBj
