[English] 日本語
 Yorodumi
Yorodumi- PDB-2k0m: Solution NMR structure of the uncharacterized protein from Rhodos... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2k0m | ||||||
|---|---|---|---|---|---|---|---|
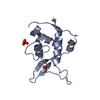
| Title | Solution NMR structure of the uncharacterized protein from Rhodospirillum rubrum gene locus Rru_A0810. Northeast Structural Genomics Target RrR43 | ||||||
 Components Components | Uncharacterized protein | ||||||
 Keywords Keywords |  STRUCTURAL GENOMICS / UNKNOWN FUNCTION / PSI-2 / STRUCTURAL GENOMICS / UNKNOWN FUNCTION / PSI-2 /  Protein Structure Initiative / Northeast Structural Genomics Consortium / NESG Protein Structure Initiative / Northeast Structural Genomics Consortium / NESG | ||||||
| Function / homology | Protein of unknown function (DUF3223) / Nuclear Transport Factor 2; Chain: A, - #40 / Nuclear Transport Factor 2; Chain: A, / Roll / Alpha Beta / Uncharacterized protein Function and homology information Function and homology information | ||||||
| Biological species |   Rhodospirillum rubrum ATCC 11170 (bacteria) Rhodospirillum rubrum ATCC 11170 (bacteria) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  molecular dynamics molecular dynamics | ||||||
 Authors Authors | Rossi, P. / Wang, H. / Jiang, M. / Foote, E.L. / Xiao, R. / Liu, J. / Swapna, G. / Acton, T.B. / Baran, M.C. / Rost, B. ...Rossi, P. / Wang, H. / Jiang, M. / Foote, E.L. / Xiao, R. / Liu, J. / Swapna, G. / Acton, T.B. / Baran, M.C. / Rost, B. / Montelione, G.T. / Northeast Structural Genomics Consortium (NESG) | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Solution NMR structure of the uncharacterized protein from Rhodospirillum rubrum gene locus Rru_A0810. Northeast Structural Genomics Target RrR43. Authors: Rossi, P. / Xiao, R. / Acton, T.B. / Montelione, G.T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2k0m.cif.gz 2k0m.cif.gz | 725.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2k0m.ent.gz pdb2k0m.ent.gz | 616.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2k0m.json.gz 2k0m.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/k0/2k0m https://data.pdbj.org/pub/pdb/validation_reports/k0/2k0m ftp://data.pdbj.org/pub/pdb/validation_reports/k0/2k0m ftp://data.pdbj.org/pub/pdb/validation_reports/k0/2k0m | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11870.565 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Rhodospirillum rubrum ATCC 11170 (bacteria) Rhodospirillum rubrum ATCC 11170 (bacteria)Species: Rhodospirillum rubrum  / Strain: NCIB 8255 / Gene: Rru_A0810 / Plasmid: pET21-23C / Production host: / Strain: NCIB 8255 / Gene: Rru_A0810 / Plasmid: pET21-23C / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3)+Magic / References: UniProt: Q2RW81 Escherichia coli (E. coli) / Strain (production host): BL21(DE3)+Magic / References: UniProt: Q2RW81 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NMR details | Text: MONOMER BY GEL FILTRATION CHROMATOGRAPHY/LIGHT SCATTERING. STRUCTURE DETERMINED BY TRIPLE RESONANCE NMR SPECTROSCOPY. NOESY ASSIGNMENTS BY CYANA2.1. 20 OF 100 STRUCTURES LOWEST TARGET FUNCTION ...Text: MONOMER BY GEL FILTRATION CHROMATOGRAPHY/LIGHT SCATTERING. STRUCTURE DETERMINED BY TRIPLE RESONANCE NMR SPECTROSCOPY. NOESY ASSIGNMENTS BY CYANA2.1. 20 OF 100 STRUCTURES LOWEST TARGET FUNCTION SELECTED. MODELS ARE FURTHER REFINED USING CNS IN EXPLICIT WATER SHELL (NILGES PROTOCOL WITH PARAM19). ASSIGNMENT STATS (EXCLUDING C-TERM TAG): BACKBONE 94.7%, SIDECHAIN 93.1%, AROMATIC (SC) 100%, VL METHYL STEREOSPECIFIC 100%, UNAMBIGUOUS SIDECHAIN NH2 100%. STRUCTURE BASED ON 1727 NOE. MAX NOE VIOLATION 0.25 A (1MODEL). 6 TOTAL CLOSE CONTACTS PER 20 MODELS. STRUCTURE QUALITY FACTOR (PSVS 1.3): ORDERED RESIDUES RANGES - 6-70, 75-94 (FOR [S(PHI)+S(PSI)] > 1.8) RMSD 0.6 BB, 1.0 ALL HEAVY ATOMS. SECONDARY STRUCTURE ELEMENTS ALPHA HELICES: (17-30, 39-50, 55-59), BETA STRANDS: (7-9, 12-14, 35-36, 63-70, 76-82, 87-89). RAMA. DISTRIBUTION: 90.2/9.7/0.1/0.1. PROCHECK (PSI-PHI): -0.29/-0.83 (RAW/Z), PROCHECK (ALL): -0.22/-1.30 (RAW/Z), MOLPROBITY CLASH: 15.46/-1.13 (RAW/Z). RPF SCORES ALL ASSIGNED RESIDUES (FIT OF NOESY PEAKLISTS TO STRUCTURE): RECALL: 0.917, PRECISION: 0.895, F-MEASURE: 0.906, DP-SCORE: 0.738. |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sample conditions | Ionic strength: 0.1 / pH: 6.5 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Bruker Avance / Manufacturer: Bruker / Model : AVANCE / Field strength: 800 MHz : AVANCE / Field strength: 800 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  molecular dynamics / Software ordinal: 1 molecular dynamics / Software ordinal: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: target function / Conformers calculated total number: 100 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller











 PDBj
PDBj