[English] 日本語
 Yorodumi
Yorodumi- PDB-2jsm: MONOMERIC HUMAN TELOMERE DNA TETRAPLEX WITH 3+1 STRAND FOLD TOPOL... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2jsm | ||||||
|---|---|---|---|---|---|---|---|
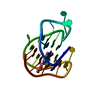
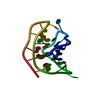
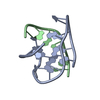
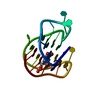
| Title | MONOMERIC HUMAN TELOMERE DNA TETRAPLEX WITH 3+1 STRAND FOLD TOPOLOGY, TWO EDGEWISE LOOPS AND DOUBLE-CHAIN REVERSAL LOOP, NMR, 10 STRUCTURES, Form 1 Natural | ||||||
 Components Components | HUMAN TELOMERE DNA | ||||||
 Keywords Keywords |  DNA / G-TETRAD / 3+1 STRAND FOLDING / DNA / G-TETRAD / 3+1 STRAND FOLDING /  EDGEWISE / DOUBLE-CHAIN REVERSAL LOOPS / Natural Form 1 / 10 STRUCTURES EDGEWISE / DOUBLE-CHAIN REVERSAL LOOPS / Natural Form 1 / 10 STRUCTURES | ||||||
| Function / homology |  DNA / DNA (> 10) DNA / DNA (> 10) Function and homology information Function and homology information | ||||||
| Method |  SOLUTION NMR / torsion angle dynamics SOLUTION NMR / torsion angle dynamics | ||||||
 Authors Authors | Kuryavyi, V.V. / Phan, A.T. / Luu, K.N. / Patel, D.J. | ||||||
 Citation Citation |  Journal: Nucleic Acids Res. / Year: 2007 Journal: Nucleic Acids Res. / Year: 2007Title: Structure of two intramolecular G-quadruplexes formed by natural human telomere sequences in K+ solution Authors: Phan, A.T. / Kuryavyi, V. / Luu, K.N. / Patel, D.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2jsm.cif.gz 2jsm.cif.gz | 144.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2jsm.ent.gz pdb2jsm.ent.gz | 119.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2jsm.json.gz 2jsm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/js/2jsm https://data.pdbj.org/pub/pdb/validation_reports/js/2jsm ftp://data.pdbj.org/pub/pdb/validation_reports/js/2jsm ftp://data.pdbj.org/pub/pdb/validation_reports/js/2jsm | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: DNA chain | Mass: 7287.690 Da / Num. of mol.: 1 / Source method: obtained synthetically |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 0.5-5.0 MM HUMAN TELOMERE DNA, 90% H2O/10% D2O |
|---|---|
| Sample | Conc.: .5 mM / Component: HUMAN TELOMERE DNA |
| Sample conditions | Ionic strength: 70 / pH: 7.0 / Pressure: 1 atm / Temperature: 298 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: torsion angle dynamics / Software ordinal: 1 Details: AFTER DEPOSITION, THE MOLECULES WERE ENERGY MINIMIZED WITH THE ENERGY FUNCTION IMPLEMENTING GEOMETRICAL VALUES IN EFFECT AT RCSB SINCE JULY 31 2007. AS A RESULT, STRUCTURE STATISTICS FOR ...Details: AFTER DEPOSITION, THE MOLECULES WERE ENERGY MINIMIZED WITH THE ENERGY FUNCTION IMPLEMENTING GEOMETRICAL VALUES IN EFFECT AT RCSB SINCE JULY 31 2007. AS A RESULT, STRUCTURE STATISTICS FOR THIS ENTRY SLIGHTLY DEVIATES FROM THE PUBLISHED ONE, AS FOLLOWS (CURRENT VS PUBLISHED): NOE VIOLATIONS (0.3+-0.48) VS (0.00+-0.00) MAXIMUM VIOLATION (0.21+-0.06) VS (0.00+-0.00) RMSD OF VIOLATIONS (0.03+-0.00) VS (0.03+-0.00) BOND LENGTHS (0.004+-0.000) VS (0.004+-0.000) BOND ANGLES (0.71+-0.01) VS (0.92+-0.01) IMPROPERS (0.38+-0.02) VS (0.36+-0.02) PAIRWISE RMSD: ALL ATOMS (0.61+-0.15) VS (0.82+-0.22) ALL ATOMS EXCEPT T6,T7,A8 (0.40+-0.11) VS (0.59+-0.15) | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller













 PDBj
PDBj






































