+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2jmo | ||||||
|---|---|---|---|---|---|---|---|
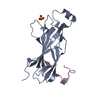
| Title | IBR domain of Human Parkin | ||||||
 Components Components | Parkin | ||||||
 Keywords Keywords |  LIGASE / Parkin / IBR / LIGASE / Parkin / IBR /  E3 ligase / zinc binding domain / RBR E3 ligase / zinc binding domain / RBR | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of primary amine oxidase activity / positive regulation of retrograde transport, endosome to Golgi / regulation of lipid transport / positive regulation of neurotransmitter uptake /  regulation protein catabolic process at presynapse / negative regulation of endoplasmic reticulum stress-induced neuron intrinsic apoptotic signaling pathway / negative regulation of spontaneous neurotransmitter secretion / regulation of protein targeting to mitochondrion / negative regulation of intralumenal vesicle formation / negative regulation of glucokinase activity ...negative regulation of primary amine oxidase activity / positive regulation of retrograde transport, endosome to Golgi / regulation of lipid transport / positive regulation of neurotransmitter uptake / regulation protein catabolic process at presynapse / negative regulation of endoplasmic reticulum stress-induced neuron intrinsic apoptotic signaling pathway / negative regulation of spontaneous neurotransmitter secretion / regulation of protein targeting to mitochondrion / negative regulation of intralumenal vesicle formation / negative regulation of glucokinase activity ...negative regulation of primary amine oxidase activity / positive regulation of retrograde transport, endosome to Golgi / regulation of lipid transport / positive regulation of neurotransmitter uptake /  regulation protein catabolic process at presynapse / negative regulation of endoplasmic reticulum stress-induced neuron intrinsic apoptotic signaling pathway / negative regulation of spontaneous neurotransmitter secretion / regulation of protein targeting to mitochondrion / negative regulation of intralumenal vesicle formation / negative regulation of glucokinase activity / negative regulation of exosomal secretion / mitochondrion to lysosome vesicle-mediated transport / positive regulation of mitochondrial fusion / parkin-mediated stimulation of mitophagy in response to mitochondrial depolarization / protein K29-linked ubiquitination / regulation protein catabolic process at presynapse / negative regulation of endoplasmic reticulum stress-induced neuron intrinsic apoptotic signaling pathway / negative regulation of spontaneous neurotransmitter secretion / regulation of protein targeting to mitochondrion / negative regulation of intralumenal vesicle formation / negative regulation of glucokinase activity / negative regulation of exosomal secretion / mitochondrion to lysosome vesicle-mediated transport / positive regulation of mitochondrial fusion / parkin-mediated stimulation of mitophagy in response to mitochondrial depolarization / protein K29-linked ubiquitination /  Lewy body / protein K27-linked ubiquitination / Parkin-FBXW7-Cul1 ubiquitin ligase complex / regulation of synaptic vesicle transport / negative regulation of mitochondrial fusion / free ubiquitin chain polymerization / negative regulation of actin filament bundle assembly / positive regulation of mitophagy in response to mitochondrial depolarization / RBR-type E3 ubiquitin transferase / positive regulation of protein linear polyubiquitination / Lewy body / protein K27-linked ubiquitination / Parkin-FBXW7-Cul1 ubiquitin ligase complex / regulation of synaptic vesicle transport / negative regulation of mitochondrial fusion / free ubiquitin chain polymerization / negative regulation of actin filament bundle assembly / positive regulation of mitophagy in response to mitochondrial depolarization / RBR-type E3 ubiquitin transferase / positive regulation of protein linear polyubiquitination /  F-box domain binding / negative regulation by host of viral genome replication / positive regulation of mitophagy / cellular response to toxic substance / regulation of dopamine metabolic process / regulation of necroptotic process / dopaminergic synapse / regulation of cellular response to oxidative stress / autophagy of mitochondrion / negative regulation of intrinsic apoptotic signaling pathway by p53 class mediator / protein K6-linked ubiquitination / positive regulation of dendrite extension / norepinephrine metabolic process / positive regulation of proteasomal protein catabolic process / protein localization to mitochondrion / negative regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway / negative regulation of JNK cascade / positive regulation of protein localization to membrane / protein K11-linked ubiquitination / cellular response to dopamine / positive regulation of tumor necrosis factor-mediated signaling pathway / F-box domain binding / negative regulation by host of viral genome replication / positive regulation of mitophagy / cellular response to toxic substance / regulation of dopamine metabolic process / regulation of necroptotic process / dopaminergic synapse / regulation of cellular response to oxidative stress / autophagy of mitochondrion / negative regulation of intrinsic apoptotic signaling pathway by p53 class mediator / protein K6-linked ubiquitination / positive regulation of dendrite extension / norepinephrine metabolic process / positive regulation of proteasomal protein catabolic process / protein localization to mitochondrion / negative regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway / negative regulation of JNK cascade / positive regulation of protein localization to membrane / protein K11-linked ubiquitination / cellular response to dopamine / positive regulation of tumor necrosis factor-mediated signaling pathway /  mitochondrial fission / mitochondrial fission /  aggresome assembly / ubiquitin conjugating enzyme binding / regulation of canonical Wnt signaling pathway / regulation of mitochondrion organization / aggresome assembly / ubiquitin conjugating enzyme binding / regulation of canonical Wnt signaling pathway / regulation of mitochondrion organization /  aggresome / regulation of reactive oxygen species metabolic process / regulation of synaptic vesicle endocytosis / dopamine uptake involved in synaptic transmission / positive regulation of mitochondrial fission / dopamine metabolic process / regulation of dopamine secretion / ubiquitin-specific protease binding / aggresome / regulation of reactive oxygen species metabolic process / regulation of synaptic vesicle endocytosis / dopamine uptake involved in synaptic transmission / positive regulation of mitochondrial fission / dopamine metabolic process / regulation of dopamine secretion / ubiquitin-specific protease binding /  startle response / ERAD pathway / negative regulation of release of cytochrome c from mitochondria / protein monoubiquitination / protein K63-linked ubiquitination / startle response / ERAD pathway / negative regulation of release of cytochrome c from mitochondria / protein monoubiquitination / protein K63-linked ubiquitination /  phospholipase binding / cullin family protein binding / phospholipase binding / cullin family protein binding /  mitophagy / regulation of protein ubiquitination / regulation of glucose metabolic process / negative regulation of insulin secretion / negative regulation of reactive oxygen species metabolic process / cellular response to unfolded protein / positive regulation of DNA binding / cellular response to manganese ion / protein autoubiquitination / protein K48-linked ubiquitination / negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway / mitophagy / regulation of protein ubiquitination / regulation of glucose metabolic process / negative regulation of insulin secretion / negative regulation of reactive oxygen species metabolic process / cellular response to unfolded protein / positive regulation of DNA binding / cellular response to manganese ion / protein autoubiquitination / protein K48-linked ubiquitination / negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway /  ubiquitin ligase complex / ubiquitin ligase complex /  Hsp70 protein binding / Hsp70 protein binding /  heat shock protein binding / PINK1-PRKN Mediated Mitophagy / response to endoplasmic reticulum stress / mitochondrion organization / heat shock protein binding / PINK1-PRKN Mediated Mitophagy / response to endoplasmic reticulum stress / mitochondrion organization /  tubulin binding / adult locomotory behavior / proteasomal protein catabolic process / Josephin domain DUBs / negative regulation of protein phosphorylation / regulation of mitochondrial membrane potential / tubulin binding / adult locomotory behavior / proteasomal protein catabolic process / Josephin domain DUBs / negative regulation of protein phosphorylation / regulation of mitochondrial membrane potential /  ubiquitin binding / ubiquitin binding /  learning / learning /  central nervous system development / central nervous system development /  synaptic transmission, glutamatergic / synaptic transmission, glutamatergic /  regulation of autophagy / G protein-coupled receptor binding / regulation of autophagy / G protein-coupled receptor binding /  PDZ domain binding / PDZ domain binding /  macroautophagy / protein destabilization / macroautophagy / protein destabilization /  regulation of protein stability / negative regulation of canonical Wnt signaling pathway regulation of protein stability / negative regulation of canonical Wnt signaling pathwaySimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing, torsion angle dynamics simulated annealing, torsion angle dynamics | ||||||
 Authors Authors | Beasley, S.A. / Hristova, V.A. / Shaw, G.S. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2007 Journal: Proc.Natl.Acad.Sci.USA / Year: 2007Title: Structure of the Parkin in-between-ring domain provides insights for E3-ligase dysfunction in autosomal recessive Parkinson's disease. Authors: Beasley, S.A. / Hristova, V.A. / Shaw, G.S. | ||||||
| History |
| ||||||
| Remark 700 | SHEET THE AUTHORS STATE THAT CD, NOE AND CHEMICAL SHIFT INDEX DATA ARE NOT CONSISTENT WITH BETA ...SHEET THE AUTHORS STATE THAT CD, NOE AND CHEMICAL SHIFT INDEX DATA ARE NOT CONSISTENT WITH BETA SHEET CHARACTER IN THIS PROTEIN. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2jmo.cif.gz 2jmo.cif.gz | 490.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2jmo.ent.gz pdb2jmo.ent.gz | 418.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2jmo.json.gz 2jmo.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jm/2jmo https://data.pdbj.org/pub/pdb/validation_reports/jm/2jmo ftp://data.pdbj.org/pub/pdb/validation_reports/jm/2jmo ftp://data.pdbj.org/pub/pdb/validation_reports/jm/2jmo | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 8618.604 Da / Num. of mol.: 1 / Fragment: IBR-type 1 domain, residues 308-384 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: PARK2, PRKN Homo sapiens (human) / Gene: PARK2, PRKNPlasmid details: the thrombin site of pET15b has been replaced with a TEV recognition site Plasmid: p11 (modified pET15b) / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3) STAR Escherichia coli (E. coli) / Strain (production host): BL21(DE3) STARReferences: UniProt: O60260,  Ligases; Forming carbon-nitrogen bonds; Acid-amino-acid ligases (peptide synthases) Ligases; Forming carbon-nitrogen bonds; Acid-amino-acid ligases (peptide synthases) |
|---|---|
| #2: Chemical |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||||||||||||||||||
| Sample conditions | Ionic strength: 100 mM NaCl, 25 mM Na2HPO4 / pH: 6.95 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| NMR spectrometer | Type: Varian INOVA / Manufacturer: Varian / Model : INOVA / Field strength: 600 MHz : INOVA / Field strength: 600 MHz |
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing, torsion angle dynamics / Software ordinal: 1 simulated annealing, torsion angle dynamics / Software ordinal: 1 Details: The authors state that they used a CYANA residue library with a modified Cysteine - zinc ligand for use in the structure calculations. These were the angles and lengths listed. | ||||||||||||||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: target function / Conformers calculated total number: 200 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller











 PDBj
PDBj



