[English] 日本語
 Yorodumi
Yorodumi- PDB-2idw: Crystal structure analysis of HIV-1 protease mutant V82A with a p... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2idw | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal structure analysis of HIV-1 protease mutant V82A with a potent non-peptide inhibitor (UIC-94017) | |||||||||
 Components Components | Protease | |||||||||
 Keywords Keywords |  HYDROLASE / HYDROLASE /  HIV-1 PROTEASE / HIV-1 PROTEASE /  MUTANT / DIMER / MUTANT / DIMER /  INHIBITOR / UIC-94017 INHIBITOR / UIC-94017 | |||||||||
| Function / homology |  Function and homology information Function and homology information: / : /  HIV-1 retropepsin / HIV-1 retropepsin /  retroviral ribonuclease H / retroviral ribonuclease H /  exoribonuclease H / exoribonuclease H /  exoribonuclease H activity / host multivesicular body / DNA integration / exoribonuclease H activity / host multivesicular body / DNA integration /  RNA-directed DNA polymerase / viral genome integration into host DNA ...: / : / RNA-directed DNA polymerase / viral genome integration into host DNA ...: / : /  HIV-1 retropepsin / HIV-1 retropepsin /  retroviral ribonuclease H / retroviral ribonuclease H /  exoribonuclease H / exoribonuclease H /  exoribonuclease H activity / host multivesicular body / DNA integration / exoribonuclease H activity / host multivesicular body / DNA integration /  RNA-directed DNA polymerase / viral genome integration into host DNA / viral penetration into host nucleus / establishment of integrated proviral latency / RNA-directed DNA polymerase / viral genome integration into host DNA / viral penetration into host nucleus / establishment of integrated proviral latency /  RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity /  Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / symbiont-mediated suppression of host gene expression / viral nucleocapsid / DNA recombination / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / symbiont-mediated suppression of host gene expression / viral nucleocapsid / DNA recombination /  Hydrolases; Acting on ester bonds / Hydrolases; Acting on ester bonds /  DNA-directed DNA polymerase / aspartic-type endopeptidase activity / DNA-directed DNA polymerase / aspartic-type endopeptidase activity /  DNA-directed DNA polymerase activity / symbiont entry into host cell / DNA-directed DNA polymerase activity / symbiont entry into host cell /  lipid binding / host cell nucleus / host cell plasma membrane / virion membrane / structural molecule activity / lipid binding / host cell nucleus / host cell plasma membrane / virion membrane / structural molecule activity /  proteolysis / proteolysis /  DNA binding / DNA binding /  RNA binding / zinc ion binding / RNA binding / zinc ion binding /  membrane / identical protein binding membrane / identical protein bindingSimilarity search - Function | |||||||||
| Biological species |    Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.1 Å MOLECULAR REPLACEMENT / Resolution: 1.1 Å | |||||||||
 Authors Authors | Tie, Y. / Boross, P.I. / Wang, Y.F. / Gaddis, L. / Manna, D. / Hussain, A.K. / Leshchenko, S. / Ghosh, A.K. / Louis, J.M. / Harrison, R.W. / Weber, I.T. | |||||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2004 Journal: J.Mol.Biol. / Year: 2004Title: High Resolution Crystal Structures of HIV-1 Protease with a Potent Non-Peptide Inhibitor (Uic-94017) Active Against Multi-Drug-Resistant Clinical Strains. Authors: Tie, Y. / Boross, P.I. / Wang, Y.F. / Gaddis, L. / Hussain, A.K. / Leshchenko, S. / Ghosh, A.K. / Louis, J.M. / Harrison, R.W. / Weber, I.T. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2idw.cif.gz 2idw.cif.gz | 114.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2idw.ent.gz pdb2idw.ent.gz | 86.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2idw.json.gz 2idw.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/id/2idw https://data.pdbj.org/pub/pdb/validation_reports/id/2idw ftp://data.pdbj.org/pub/pdb/validation_reports/id/2idw ftp://data.pdbj.org/pub/pdb/validation_reports/id/2idw | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 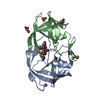 2ienC  2ieoC  1dazS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 2 molecules AB
| #1: Protein |  / RETROPEPSIN / RETROPEPSINMass: 10712.623 Da / Num. of mol.: 2 / Fragment: residues 500-598 Mutation: Q7K, L33I, L63I, C67A, V82A, C95A, Q107K, L133I, L163I, C167A, V182A, C195A Source method: isolated from a genetically manipulated source Source: (gene. exp.)    Human immunodeficiency virus 1 / Genus: Lentivirus Human immunodeficiency virus 1 / Genus: Lentivirus / Gene: gag, pol / Plasmid: PET11A / Species (production host): Escherichia coli / Production host: / Gene: gag, pol / Plasmid: PET11A / Species (production host): Escherichia coli / Production host:   Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21(DE3) Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21(DE3)References: UniProt: P03368, UniProt: Q7SSI0*PLUS,  HIV-1 retropepsin HIV-1 retropepsin |
|---|
-Non-polymers , 7 types, 271 molecules 











| #2: Chemical | ChemComp-CL /  Chloride Chloride | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| #3: Chemical | ChemComp-PO4 /  Phosphate Phosphate#4: Chemical | ChemComp-DMS / |  Dimethyl sulfoxide Dimethyl sulfoxide#5: Chemical | ChemComp-ACY /  Acetic acid Acetic acid#6: Chemical | ChemComp-GOL / |  Glycerol Glycerol#7: Chemical | ChemComp-017 / ( |  Darunavir Darunavir#8: Water | ChemComp-HOH / |  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.07 Å3/Da / Density % sol: 40.08 % |
|---|---|
Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 5 Details: AMMONIUM SULFATE, CITRIC PHOSPHATE, DMSO, pH 5.00, VAPOR DIFFUSION, HANGING DROP, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 90 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 1 / Beamline: 22-ID / Wavelength: 1 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Dec 10, 2002 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.1→50 Å / Num. obs: 74707 / % possible obs: 99.7 % / Observed criterion σ(I): 3 / Redundancy: 5.4 % / Rmerge(I) obs: 0.052 / Net I/σ(I): 18.4 |
| Reflection shell | Resolution: 1.1→1.14 Å / Redundancy: 2.4 % / Rmerge(I) obs: 0.196 / % possible all: 97.6 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1DAZ Resolution: 1.1→10 Å / Num. parameters: 25871 / Num. restraintsaints: 24400 / σ(F): 0 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||||||||||
| Refine analyze | Num. disordered residues: 43 / Occupancy sum hydrogen: 1624 / Occupancy sum non hydrogen: 1768 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.1→10 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj









