+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2cbp | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CUCUMBER BASIC PROTEIN, A BLUE COPPER PROTEIN | |||||||||
 Components Components | CUCUMBER BASIC PROTEIN | |||||||||
 Keywords Keywords |  ELECTRON TRANSPORT / PHYTOCYANIN / TYPE 1 COPPER PROTEIN ELECTRON TRANSPORT / PHYTOCYANIN / TYPE 1 COPPER PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information pollination / anther development / pollination / anther development /  apoplast / apoplast /  extracellular matrix / extracellular matrix /  electron transfer activity / electron transfer activity /  metal ion binding metal ion bindingSimilarity search - Function | |||||||||
| Biological species |   Cucumis sativus (cucumber) Cucumis sativus (cucumber) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 1.8 Å SYNCHROTRON / Resolution: 1.8 Å | |||||||||
 Authors Authors | Guss, J.M. / Freeman, H.C. | |||||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: The structure of a phytocyanin, the basic blue protein from cucumber, refined at 1.8 A resolution. Authors: Guss, J.M. / Merritt, E.A. / Phizackerley, R.P. / Freeman, H.C. #1:  Journal: Science / Year: 1988 Journal: Science / Year: 1988Title: Phase Determination by Multiple-Wavelength X-Ray Diffraction. Crystal Structure of a Basic 'Blue' Copper Protein from Cucumbers Authors: Guss, J.M. / Merritt, E.A. / Phizackerley, R.P. / Hedman, B. / Murata, M. / Hodgson, K.O. / Freeman, H.C. #2:  Journal: J.Mol.Biol. / Year: 1977 Journal: J.Mol.Biol. / Year: 1977Title: Preliminary Crystallographic Data for a Basic Copper-Containing Protein from Cucumber Seedlings Authors: Colman, P.M. / Freeman, H.C. / Guss, J.M. / Murata, M. / Norris, V.A. / Ramshaw, J.A. / Venkatappa, M.P. / Vickery, L.E. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2cbp.cif.gz 2cbp.cif.gz | 51.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2cbp.ent.gz pdb2cbp.ent.gz | 37.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2cbp.json.gz 2cbp.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/cb/2cbp https://data.pdbj.org/pub/pdb/validation_reports/cb/2cbp ftp://data.pdbj.org/pub/pdb/validation_reports/cb/2cbp ftp://data.pdbj.org/pub/pdb/validation_reports/cb/2cbp | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 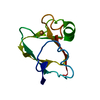
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 10371.791 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Details: FROM SEEDLINGS / Source: (natural)   Cucumis sativus (cucumber) / References: UniProt: P00303 Cucumis sativus (cucumber) / References: UniProt: P00303 |
|---|---|
| #2: Chemical | ChemComp-CU /  Copper Copper |
| #3: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.26 Å3/Da / Density % sol: 47 % | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | *PLUS pH: 6 / Method: vapor diffusion, hanging drop | |||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL1-5 / Wavelength: 1.2359, 1.3771, 1.3790, 1.5416 / Beamline: BL1-5 / Wavelength: 1.2359, 1.3771, 1.3790, 1.5416 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Type: XUONG-HAMLIN MULTIWIRE / Detector: AREA DETECTOR / Date: 1986 | |||||||||||||||
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||
| Radiation wavelength |
| |||||||||||||||
| Reflection | Num. obs: 43466 / % possible obs: 89 % / Observed criterion σ(I): 1 / Rmerge(I) obs: 0.043 |
- Processing
Processing
| Software | Name: PROLSQ / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.8→7 Å / σ(F): 0.5 Details: 1988 ESTIMATED COORD. ERROR 0.12 ANGSTROMS (BACKBONE) ESTIMATED COORD. ERROR 0.17 ANGSTROMS (SIDE CHAIN)
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 11.9 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→7 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.141 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller











 PDBj
PDBj


