+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2bnh | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PORCINE RIBONUCLEASE INHIBITOR | |||||||||
 Components Components | RIBONUCLEASE INHIBITOR | |||||||||
 Keywords Keywords |  ACETYLATION / ACETYLATION /  LEUCINE-RICH REPEATS LEUCINE-RICH REPEATS | |||||||||
| Function / homology |  Function and homology information Function and homology information ribonuclease inhibitor activity / angiogenin-PRI complex / regulation of Arp2/3 complex-mediated actin nucleation / ribonuclease inhibitor activity / angiogenin-PRI complex / regulation of Arp2/3 complex-mediated actin nucleation /  regulation of angiogenesis / regulation of angiogenesis /  cell migration / cell migration /  lamellipodium / lamellipodium /  regulation of inflammatory response / regulation of inflammatory response /  nucleoplasm / nucleoplasm /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Sus scrofa (pig) Sus scrofa (pig) | |||||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.3 Å X-RAY DIFFRACTION / Resolution: 2.3 Å | |||||||||
 Authors Authors | Kobe, B. / Deisenhofer, J. | |||||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: Mechanism of ribonuclease inhibition by ribonuclease inhibitor protein based on the crystal structure of its complex with ribonuclease A. Authors: Kobe, B. / Deisenhofer, J. #1:  Journal: J.Mol.Biol. / Year: 1993 Journal: J.Mol.Biol. / Year: 1993Title: Crystallization and Preliminary X-Ray Analysis of Porcine Ribonuclease Inhibitor, a Protein with Leucine-Rich Repeats Authors: Kobe, B. / Deisenhofer, J. #2:  Journal: Nature / Year: 1993 Journal: Nature / Year: 1993Title: Crystal Structure of Porcine Ribonuclease Inhibitor, a Protein with Leucine-Rich Repeats Authors: Kobe, B. / Deisenhofer, J. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2bnh.cif.gz 2bnh.cif.gz | 96.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2bnh.ent.gz pdb2bnh.ent.gz | 73.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2bnh.json.gz 2bnh.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bn/2bnh https://data.pdbj.org/pub/pdb/validation_reports/bn/2bnh ftp://data.pdbj.org/pub/pdb/validation_reports/bn/2bnh ftp://data.pdbj.org/pub/pdb/validation_reports/bn/2bnh | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 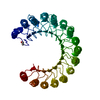
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  / RNASE INHIBITOR / RIBONUCLEASE/ANGIOGENIN INHIBITOR / RNASE INHIBITOR / RIBONUCLEASE/ANGIOGENIN INHIBITORMass: 49092.156 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Sus scrofa (pig) / Organ: LIVER Sus scrofa (pig) / Organ: LIVER / References: UniProt: P10775 / References: UniProt: P10775 |
|---|---|
| #2: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.87 Å3/Da / Density % sol: 68.24 % | ||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | *PLUS Temperature: 21 ℃ / pH: 7 / Method: unknown | ||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Wavelength: 1.5418 |
|---|---|
| Detector | Type: XUONG-HAMLIN MULTIWIRE / Detector: AREA DETECTOR / Date: 1992 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 |
| Reflection | Num. obs: 24468 / % possible obs: 73.2 % / Observed criterion σ(I): 1 / Redundancy: 5.7 % / Rmerge(I) obs: 0.071 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2.3→40 Å / σ(F): 1 /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 30.4 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.32 Å / Luzzati sigma a obs: 0.49 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→40 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller











 PDBj
PDBj








