+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1tof | ||||||
|---|---|---|---|---|---|---|---|
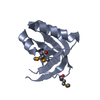
| Title | THIOREDOXIN H (OXIDIZED FORM), NMR, 23 STRUCTURES | ||||||
 Components Components | THIOREDOXIN H | ||||||
 Keywords Keywords |  ELECTRON TRANSPORT / ELECTRON TRANSPORT /  OXIDOREDUCTASE OXIDOREDUCTASE | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |   Chlamydomonas reinhardtii (plant) Chlamydomonas reinhardtii (plant) | ||||||
| Method |  SOLUTION NMR SOLUTION NMR | ||||||
 Authors Authors | Mittard, V. / Blackledge, M.J. / Stein, M. / Jacquot, J.-P. / Marion, D. / Lancelin, J.-M. | ||||||
 Citation Citation |  Journal: Eur.J.Biochem. / Year: 1997 Journal: Eur.J.Biochem. / Year: 1997Title: NMR solution structure of an oxidised thioredoxin h from the eukaryotic green alga Chlamydomonas reinhardtii. Authors: Mittard, V. / Blackledge, M.J. / Stein, M. / Jacquot, J.P. / Marion, D. / Lancelin, J.M. #1:  Journal: Plant Mol.Biol. / Year: 1995 Journal: Plant Mol.Biol. / Year: 1995Title: Chlamydomonas Reinhardtii Thioredoxins: Structure of the Genes Coding for the Chloroplastic M and Cytosolic H Isoforms; Expression in Escherichia Coli of the Recombinant Proteins, Purification ...Title: Chlamydomonas Reinhardtii Thioredoxins: Structure of the Genes Coding for the Chloroplastic M and Cytosolic H Isoforms; Expression in Escherichia Coli of the Recombinant Proteins, Purification and Biochemical Properties Authors: Stein, M. / Jacquot, J.P. / Jeannette, E. / Decottignies, P. / Hodges, M. / Lancelin, J.M. / Mittard, V. / Schmitter, J.M. / Miginiac-Maslow, M. #2:  Journal: Eur.J.Biochem. / Year: 1995 Journal: Eur.J.Biochem. / Year: 1995Title: 1H, 13C, 15N-NMR Resonance Assignments of Oxidized Thioredoxin H from the Eukaryotic Green Alga Chlamydomonas Reinhardtii Using New Methods Based on Two-Dimensional Triple-Resonance NMR ...Title: 1H, 13C, 15N-NMR Resonance Assignments of Oxidized Thioredoxin H from the Eukaryotic Green Alga Chlamydomonas Reinhardtii Using New Methods Based on Two-Dimensional Triple-Resonance NMR Spectroscopy and Computer-Assisted Backbone Assignment Authors: Mittard, V. / Morelle, N. / Brutscher, B. / Simorre, J.P. / Marion, D. / Stein, M. / Jacquot, J.P. / Lirsac, P.N. / Lancelin, J.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1tof.cif.gz 1tof.cif.gz | 734.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1tof.ent.gz pdb1tof.ent.gz | 638.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1tof.json.gz 1tof.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/to/1tof https://data.pdbj.org/pub/pdb/validation_reports/to/1tof ftp://data.pdbj.org/pub/pdb/validation_reports/to/1tof ftp://data.pdbj.org/pub/pdb/validation_reports/to/1tof | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11727.534 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Chlamydomonas reinhardtii (plant) / Strain: CW15 / Cellular location: CYTOPLASM Chlamydomonas reinhardtii (plant) / Strain: CW15 / Cellular location: CYTOPLASM / Gene: THIOREDOXIN H CDNA (X78822 EMB / Plasmid: PET-3D / Species (production host): Escherichia coli / Gene (production host): THIOREDOXIN H CDNA (X78822,EMBL) / Production host: / Gene: THIOREDOXIN H CDNA (X78822 EMB / Plasmid: PET-3D / Species (production host): Escherichia coli / Gene (production host): THIOREDOXIN H CDNA (X78822,EMBL) / Production host:   Escherichia coli BL21 (bacteria) / Strain (production host): BL21 / References: UniProt: P80028 Escherichia coli BL21 (bacteria) / Strain (production host): BL21 / References: UniProt: P80028 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
Crystal grow | *PLUS Method: other / Details: NMR |
|---|
- Processing
Processing
| NMR software | Name: Discover / Developer: BIOSYM / Classification: refinement |
|---|---|
| NMR ensemble | Conformers submitted total number: 23 |
 Movie
Movie Controller
Controller












 PDBj
PDBj

