[English] 日本語
 Yorodumi
Yorodumi- PDB-1pu9: Crystal Structure of Tetrahymena GCN5 with Bound Coenzyme A and a... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1pu9 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of Tetrahymena GCN5 with Bound Coenzyme A and a 19-residue Histone H3 Peptide | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSFERASE/STRUCTURAL PROTEIN /  HISTONE ACETYLTRANSFERASE / GCN5-RELATED N-ACETYLTRANSFERASE / COA-BINDING PROTEIN / HISTONE ACETYLTRANSFERASE / GCN5-RELATED N-ACETYLTRANSFERASE / COA-BINDING PROTEIN /  TERNARY COMPLEX / TRANSFERASE-STRUCTURAL PROTEIN COMPLEX TERNARY COMPLEX / TRANSFERASE-STRUCTURAL PROTEIN COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationSIRT1 negatively regulates rRNA expression / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / Oxidative Stress Induced Senescence / chromatin organization => GO:0006325 / Assembly of the ORC complex at the origin of replication / sexual sporulation resulting in formation of a cellular spore / Condensation of Prophase Chromosomes / global genome nucleotide-excision repair / RNA polymerase I upstream activating factor complex / replication fork protection complex ...SIRT1 negatively regulates rRNA expression / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / Oxidative Stress Induced Senescence / chromatin organization => GO:0006325 / Assembly of the ORC complex at the origin of replication / sexual sporulation resulting in formation of a cellular spore / Condensation of Prophase Chromosomes / global genome nucleotide-excision repair / RNA polymerase I upstream activating factor complex / replication fork protection complex / RNA Polymerase I Promoter Escape / Estrogen-dependent gene expression / positive regulation of transcription by RNA polymerase I / nucleolar large rRNA transcription by RNA polymerase I / rRNA transcription / CENP-A containing nucleosome /  histone acetyltransferase activity / histone acetyltransferase activity /  histone acetyltransferase / structural constituent of chromatin / histone acetyltransferase / structural constituent of chromatin /  nucleosome / chromatin organization / protein heterodimerization activity / regulation of DNA-templated transcription / nucleosome / chromatin organization / protein heterodimerization activity / regulation of DNA-templated transcription /  DNA binding / DNA binding /  nucleus nucleusSimilarity search - Function | ||||||
| Biological species |   Tetrahymena thermophila (eukaryote) Tetrahymena thermophila (eukaryote) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.3 Å MOLECULAR REPLACEMENT / Resolution: 2.3 Å | ||||||
 Authors Authors | Clements, A. / Poux, A.N. / Lo, W.S. / Pillus, L. / Berger, S.L. / Marmorstein, R. | ||||||
 Citation Citation |  Journal: Mol.Cell / Year: 2003 Journal: Mol.Cell / Year: 2003Title: Structural basis for histone and phospho-histone binding by the GCN5 histone acetyltransferase Authors: Clements, A. / Poux, A.N. / Lo, W.S. / Pillus, L. / Berger, S.L. / Marmorstein, R. | ||||||
| History |
| ||||||
| Remark 999 | SEQUENCE The author state: " This construct was cloned directly from the Tetrahymena thermophilia ...SEQUENCE The author state: " This construct was cloned directly from the Tetrahymena thermophilia genome. It is assumed that the protein sequence is correct, particularly in the case of the phenylalanine, as it is highly conserved among Gcn5 histone acetyltransferases from other species." |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1pu9.cif.gz 1pu9.cif.gz | 55.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1pu9.ent.gz pdb1pu9.ent.gz | 38.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1pu9.json.gz 1pu9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pu/1pu9 https://data.pdbj.org/pub/pdb/validation_reports/pu/1pu9 ftp://data.pdbj.org/pub/pdb/validation_reports/pu/1pu9 ftp://data.pdbj.org/pub/pdb/validation_reports/pu/1pu9 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1puaC  1q2cC 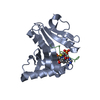 1qsnS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 19457.658 Da / Num. of mol.: 1 / Fragment: residues 48-210 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Tetrahymena thermophila (eukaryote) / Plasmid: PRSETA / Species (production host): Escherichia coli / Production host: Tetrahymena thermophila (eukaryote) / Plasmid: PRSETA / Species (production host): Escherichia coli / Production host:   Escherichia coli BL21 (bacteria) / Strain (production host): BL-21 / References: UniProt: Q27198 Escherichia coli BL21 (bacteria) / Strain (production host): BL-21 / References: UniProt: Q27198 |
|---|---|
| #2: Protein/peptide |  Mass: 2019.352 Da / Num. of mol.: 1 / Fragment: Residues 305-323 / Source method: obtained synthetically Details: This protein naturally occurs in Saccharomyces cerevisiae (Baker's yeast). References: UniProt: P02303, UniProt: P61830*PLUS |
| #3: Chemical | ChemComp-COA /  Coenzyme A Coenzyme A |
| #4: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.71 Å3/Da / Density % sol: 54.68 % | |||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 298 K / pH: 7.5 Details: Ammonium sulfate, MnCl2, Tris-HCl, pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 298K, pH 7.50 | |||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / pH: 7.5 / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CHESS CHESS  / Beamline: A1 / Wavelength: 0.9213 / Beamline: A1 / Wavelength: 0.9213 |
|---|---|
| Detector | Type: ADSC QUANTUM 210 / Detector: CCD / Date: Jun 8, 2000 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.9213 Å / Relative weight: 1 : 0.9213 Å / Relative weight: 1 |
| Reflection | Resolution: 2.24→27 Å / Num. obs: 10007 / % possible obs: 96 % / Observed criterion σ(I): 0 / Biso Wilson estimate: 20.4 Å2 |
| Reflection shell | Resolution: 2.24→2.39 Å |
| Reflection | *PLUS Highest resolution: 2.21 Å / Lowest resolution: 50 Å / Num. obs: 22051 / % possible obs: 99.5 % / Num. measured all: 67990 / Rmerge(I) obs: 0.092 |
| Reflection shell | *PLUS Highest resolution: 2.21 Å / Lowest resolution: 2.35 Å / % possible obs: 99.5 % / Rmerge(I) obs: 0.18 / Mean I/σ(I) obs: 1.3 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1QSN Resolution: 2.3→21.12 Å / Rfactor Rfree error: 0.008 / Data cutoff high absF: 1492487.31 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 61.19 Å2 / ksol: 0.45 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 27.2 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→21.12 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.3→2.44 Å / Rfactor Rfree error: 0.022 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 50 Å / % reflection Rfree: 10 % / Rfactor Rfree : 0.277 / Rfactor Rwork : 0.277 / Rfactor Rwork : 0.232 : 0.232 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller















 PDBj
PDBj















