+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1lpj | ||||||
|---|---|---|---|---|---|---|---|
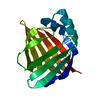
| Title | Human cRBP IV | ||||||
 Components Components | Retinol-binding protein IV, cellular | ||||||
 Keywords Keywords |  TRANSPORT PROTEIN / cellular retinol-binding protein / cRBP / TRANSPORT PROTEIN / cellular retinol-binding protein / cRBP /  retinol / vitamin A / iLBPs retinol / vitamin A / iLBPs | ||||||
| Function / homology |  Function and homology information Function and homology information retinal binding / retinal binding /  retinol binding / fatty acid transport / retinol binding / fatty acid transport /  fatty acid binding / fatty acid binding /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Calderone, V. / Zanotti, G. / Berni, R. / Folli, C. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2002 Journal: J.Biol.Chem. / Year: 2002Title: Ligand binding and structural analysis of a human putative cellular retinol-binding protein Authors: Folli, C. / Calderone, V. / Ramazzina, I. / Zanotti, G. / Berni, R. #1:  Journal: J.Mol.Biol. / Year: 1993 Journal: J.Mol.Biol. / Year: 1993Title: Crystallographic studies on a family of cellular lipophilic transport proteins. Refinement of P2 myelin protein and the structure determination and refinement of cellular retinol-binding ...Title: Crystallographic studies on a family of cellular lipophilic transport proteins. Refinement of P2 myelin protein and the structure determination and refinement of cellular retinol-binding protein in complex with all-trans-retinol. Authors: Cowan, S.W. / Newcomer, M.E. / Jones, T.A. #2:  Journal: J.Mol.Biol. / Year: 1993 Journal: J.Mol.Biol. / Year: 1993Title: Crystal structures of holo and apo-cellular retinol-binding protein II Authors: Winter, N.S. / Bratt, J.M. / Banaszak, L.J. #3:  Journal: Proc.Natl.Acad.Sci.USA / Year: 2001 Journal: Proc.Natl.Acad.Sci.USA / Year: 2001Title: Identification, Retinoid Binding and X-Ray Analysis of a Human Retinol-Binding Protein Authors: Folli, C. / Calderone, V. / Ottonello, S. / Bolchi, A. / Zanotti, G. / Stoppini, M. / Berni, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1lpj.cif.gz 1lpj.cif.gz | 43.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1lpj.ent.gz pdb1lpj.ent.gz | 29.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1lpj.json.gz 1lpj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lp/1lpj https://data.pdbj.org/pub/pdb/validation_reports/lp/1lpj ftp://data.pdbj.org/pub/pdb/validation_reports/lp/1lpj ftp://data.pdbj.org/pub/pdb/validation_reports/lp/1lpj | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1crbS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 15427.511 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) / References: UniProt: Q96R05 Escherichia coli (E. coli) / References: UniProt: Q96R05 |
|---|---|
| #2: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.08 Å3/Da / Density % sol: 40.86 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 5.6 Details: 10% PEG 4000, 10% isopropanol, 100 mM sodium citrate, pH 5.6, VAPOR DIFFUSION, SITTING DROP, temperature 293K | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 18 ℃ | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID29 / Wavelength: 0.9393 Å / Beamline: ID29 / Wavelength: 0.9393 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Feb 14, 2002 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.9393 Å / Relative weight: 1 : 0.9393 Å / Relative weight: 1 |
| Reflection | Resolution: 2→28.606 Å / Num. all: 7611 / Num. obs: 7611 / % possible obs: 89.6 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 2.7 % / Biso Wilson estimate: 23.2 Å2 / Rmerge(I) obs: 0.124 / Rsym value: 0.124 / Net I/σ(I): 3.8 |
| Reflection shell | Resolution: 2→2.1 Å / Redundancy: 2.7 % / Rmerge(I) obs: 0.382 / Mean I/σ(I) obs: 1.3 / Num. unique all: 932 / Rsym value: 0.124 / % possible all: 87.8 |
| Reflection | *PLUS Lowest resolution: 28.6 Å / Num. measured all: 19128 |
| Reflection shell | *PLUS Lowest resolution: 2.11 Å / % possible obs: 91.4 % |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1CRB Resolution: 2→24.44 Å / Rfactor Rfree error: 0.01 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 112.525 Å2 / ksol: 0.422063 e/Å3 | |||||||||||||||||||||||||
| Displacement parameters | Biso mean: 36.1 Å2
| |||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error free: 0.35 Å / Luzzati sigma a free: 0.17 Å | |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→24.44 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.13 Å / Rfactor Rfree error: 0.026 / Total num. of bins used: 6
| |||||||||||||||||||||||||
| Xplor file |
| |||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree : 0.276 / Rfactor Rwork : 0.276 / Rfactor Rwork : 0.234 : 0.234 | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| |||||||||||||||||||||||||
| LS refinement shell | *PLUS Highest resolution: 2 Å / Lowest resolution: 2.11 Å / Rfactor Rfree: 0.3 / Num. reflection Rfree: 136 |
 Movie
Movie Controller
Controller















 PDBj
PDBj


