[English] 日本語
 Yorodumi
Yorodumi- PDB-1kos: SOLUTION NMR STRUCTURE OF AN ANALOG OF THE YEAST TRNA PHE T STEM ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1kos | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | SOLUTION NMR STRUCTURE OF AN ANALOG OF THE YEAST TRNA PHE T STEM LOOP CONTAINING RIBOTHYMIDINE AT ITS NATURALLY OCCURRING POSITION | ||||||||||||||||||
 Components Components | 5'-R(* Keywords Keywords RNA / RNA /  TRNA / T STEM LOOP / TRNA / T STEM LOOP /  TRNA DOMAIN / RNA HAIRPIN / RNA FOLDING TRNA DOMAIN / RNA HAIRPIN / RNA FOLDINGFunction / homology |  RNA / RNA (> 10) RNA / RNA (> 10) Function and homology information Function and homology informationMethod |  SOLUTION NMR / DISTANCE GEOMETRY, SOLUTION NMR / DISTANCE GEOMETRY,  SIMULATED ANNEALING SIMULATED ANNEALING  Authors AuthorsKoshlap, K.M. / Guenther, R. / Sochacka, E. / Malkiewicz, A. / Agris, P.F. |  Citation Citation Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: A distinctive RNA fold: the solution structure of an analogue of the yeast tRNAPhe T Psi C domain. Authors: Koshlap, K.M. / Guenther, R. / Sochacka, E. / Malkiewicz, A. / Agris, P.F. #1:  Journal: Biopolymers / Year: 1998 Journal: Biopolymers / Year: 1998Title: Small Structural Ensembles for a 17-Nucleotide Mimic of the tRNA Tpsic-Loop Via Fitting Dipolar Relaxation Rates with the Quadratic Programming Algorithm Authors: Schmitz, U. / Donati, A. / James, T.L. / Ulyanov, N.B. / Yao, L. #2:  Journal: J.Biomol.NMR / Year: 1997 Journal: J.Biomol.NMR / Year: 1997Title: The Dynamic NMR Structure of the T Psi C-Loop: Implications for the Specificity of tRNA Methylation Authors: Yao, L.J. / James, T.L. / Kealey, J.T. / Santi, D.V. / Schmitz, U. #3:  Journal: Biochemistry / Year: 1986 Journal: Biochemistry / Year: 1986Title: Restrained Refinement of the Monoclinic Form of Yeast Phenylalanine Transfer RNA. Temperature Factors and Dynamics, Coordinated Waters, and Base-Pair Propeller Twist Angles Authors: Westhof, E. / Sundaralingam, M. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1kos.cif.gz 1kos.cif.gz | 98.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1kos.ent.gz pdb1kos.ent.gz | 74.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1kos.json.gz 1kos.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ko/1kos https://data.pdbj.org/pub/pdb/validation_reports/ko/1kos ftp://data.pdbj.org/pub/pdb/validation_reports/ko/1kos ftp://data.pdbj.org/pub/pdb/validation_reports/ko/1kos | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 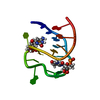
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: RNA chain | Mass: 5395.255 Da / Num. of mol.: 1 / Fragment: TPSIC DOMAIN OF TRNA / Mutation: +C49C / Source method: obtained synthetically Details: SEQUENCE FROM YEAST (SACCHAROMYCES CEREVISIAE) TRNA PHE |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||
| NMR details | Text: EXPERIMENTS IN 100% D2O CONDUCTED AT 283, 288,293, AND 298 KELVIN; EXPERIMENTS IN 6% D2O/ 94% H2O CONDUCTED AT 274 AND 283 KELVIN. |
- Sample preparation
Sample preparation
| Details | Contents: 100% D2O, 94% WATER/6% D2O |
|---|---|
| Sample conditions | pH: 6.0 / Temperature: 298 K |
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker DRX500 / Manufacturer: Bruker / Model : DRX500 / Field strength: 500 MHz : DRX500 / Field strength: 500 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: DISTANCE GEOMETRY,  SIMULATED ANNEALING / Software ordinal: 1 SIMULATED ANNEALING / Software ordinal: 1 Details: REFINEMENT DETAILS CAN BE FOUND IN THE JRNL CITATION ABOVE | ||||||||||||
| NMR ensemble | Conformer selection criteria: LOWEST ENERGY STRUCTURES / Conformers calculated total number: 50 / Conformers submitted total number: 8 |
 Movie
Movie Controller
Controller








 PDBj
PDBj





























