[English] 日本語
 Yorodumi
Yorodumi- PDB-1i8k: CRYSTAL STRUCTURE OF DSFV MR1 IN COMPLEX WITH THE PEPTIDE ANTIGEN... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1i8k | ||||||
|---|---|---|---|---|---|---|---|
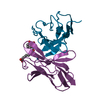
| Title | CRYSTAL STRUCTURE OF DSFV MR1 IN COMPLEX WITH THE PEPTIDE ANTIGEN OF THE MUTANT EPIDERMAL GROWTH FACTOR RECEPTOR, EGFRVIII, AT LIQUID NITROGEN TEMPERATURE | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  IMMUNE SYSTEM / antibody-peptide complex / immunoglobulin fold / peptide antigen / type II' beta turn. IMMUNE SYSTEM / antibody-peptide complex / immunoglobulin fold / peptide antigen / type II' beta turn. | ||||||
| Function / homology |  Function and homology information Function and homology information immunoglobulin complex / immunoglobulin mediated immune response / immunoglobulin complex / immunoglobulin mediated immune response /  antigen binding / antigen binding /  adaptive immune response / adaptive immune response /  immune response / immune response /  extracellular space extracellular spaceSimilarity search - Function | ||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | ||||||
 Authors Authors | Landry, R.C. / Klimowicz, A.C. / Lavictoire, S.J. / Borisova, S. / Kottachchi, D.T. / Lorimer, I.A. / Evans, S.V. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2001 Journal: J.Mol.Biol. / Year: 2001Title: Antibody recognition of a conformational epitope in a peptide antigen: Fv-peptide complex of an antibody fragment specific for the mutant EGF receptor, EGFRvIII. Authors: Landry, R.C. / Klimowicz, A.C. / Lavictoire, S.J. / Borisova, S. / Kottachchi, D.T. / Lorimer, I.A. / Evans, S.V. | ||||||
| History |
| ||||||
| Remark 999 | SEQUENCE Residues 100 of chain A and 344 of chain B have been mutated to Cys residues to introduce ...SEQUENCE Residues 100 of chain A and 344 of chain B have been mutated to Cys residues to introduce a disulfide bridge between chains A and B. There is no sequence database match for chain C because the sequence of EGFRvIII has not yet been deposited in any database. The DNA sequence of the normal EGF receptor has been deposited (NM_005228). |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1i8k.cif.gz 1i8k.cif.gz | 60.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1i8k.ent.gz pdb1i8k.ent.gz | 47.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1i8k.json.gz 1i8k.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/i8/1i8k https://data.pdbj.org/pub/pdb/validation_reports/i8/1i8k ftp://data.pdbj.org/pub/pdb/validation_reports/i8/1i8k ftp://data.pdbj.org/pub/pdb/validation_reports/i8/1i8k | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 11745.038 Da / Num. of mol.: 1 / Mutation: D100C Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Production host: Mus musculus (house mouse) / Production host:   Escherichia coli (E. coli) / Strain (production host): TG1 / References: UniProt: Q8R028, UniProt: P01660*PLUS Escherichia coli (E. coli) / Strain (production host): TG1 / References: UniProt: Q8R028, UniProt: P01660*PLUS |
|---|---|
| #2: Protein | Mass: 13712.354 Da / Num. of mol.: 1 / Mutation: R344C Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Production host: Mus musculus (house mouse) / Production host:   Escherichia coli (E. coli) / Strain (production host): TG1 / References: UniProt: P18529 Escherichia coli (E. coli) / Strain (production host): TG1 / References: UniProt: P18529 |
| #3: Protein/peptide |  / EPIDERMAL GROWTH FACTOR RECEPTOR / EGFRVIII PEPTIDE ANTIGEN / EPIDERMAL GROWTH FACTOR RECEPTOR / EGFRVIII PEPTIDE ANTIGENMass: 1421.531 Da / Num. of mol.: 1 / Fragment: N-TERMINAL FRAGMENT / Source method: obtained synthetically Details: The peptide sequence is from the N-terminal of the mutant epidermal growth factor receptor, EGFRvIII |
| #4: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.5 Å3/Da / Density % sol: 50.8 % | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 7 Details: sodium chloride, MPD, PEG 10K, pH 7.0, VAPOR DIFFUSION, HANGING DROP, temperature 293K | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 18 ℃ | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X8C / Wavelength: 0.9789 Å / Beamline: X8C / Wavelength: 0.9789 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Jun 28, 1999 |
| Radiation | Monochromator: collimating mirror optics / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.9789 Å / Relative weight: 1 : 0.9789 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→6 Å / Num. all: 25057 / Num. obs: 24309 / % possible obs: 99.8 % / Observed criterion σ(F): 3 / Observed criterion σ(I): 3 / Redundancy: 7.26 % / Biso Wilson estimate: 13.94 Å2 / Rmerge(I) obs: 0.052 / Net I/σ(I): 18.6 |
| Reflection shell | Resolution: 1.8→1.86 Å / Redundancy: 6.94 % / Rmerge(I) obs: 0.143 / % possible all: 99.4 |
| Reflection | *PLUS Highest resolution: 1.8 Å / Num. obs: 24641 / % possible obs: 98.5 % / Num. measured all: 25654 |
| Reflection shell | *PLUS % possible obs: 95.7 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT / Resolution: 1.8→6 Å / σ(F): 3 / σ(I): 3 / Stereochemistry target values: CHARMm MOLECULAR REPLACEMENT / Resolution: 1.8→6 Å / σ(F): 3 / σ(I): 3 / Stereochemistry target values: CHARMm
| ||||||||||||||||||||
| Displacement parameters | Biso mean: 13 Å2 | ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→6 Å
| ||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||
| LS refinement shell | Resolution: 1.8→1.88 Å
| ||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.8 Å / Lowest resolution: 6 Å / σ(F): 3 / Rfactor Rfree : 0.225 : 0.225 | ||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 13 Å2 | ||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.276 / Rfactor Rwork: 0.216 |
 Movie
Movie Controller
Controller












 PDBj
PDBj
