[English] 日本語
 Yorodumi
Yorodumi- PDB-1emy: CRYSTAL STRUCTURE OF ASIAN ELEPHANT (ELEPHAS MAXIMUS) CYANO-MET M... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1emy | ||||||
|---|---|---|---|---|---|---|---|
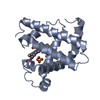
| Title | CRYSTAL STRUCTURE OF ASIAN ELEPHANT (ELEPHAS MAXIMUS) CYANO-MET MYOGLOBIN AT 1.78 ANGSTROMS RESOLUTION. PHE 29 (B10) ACCOUNTS FOR ITS UNUSUAL LIGAND BINDING PROPERTIES | ||||||
 Components Components | MYOGLOBIN | ||||||
 Keywords Keywords |  OXYGEN TRANSPORT / OXYGEN TRANSPORT /  HEME PROTEIN / HEME PROTEIN /  GLOBIN FOLD GLOBIN FOLD | ||||||
| Function / homology |  Function and homology information Function and homology information oxygen carrier activity / oxygen carrier activity /  oxygen binding / oxygen binding /  heme binding / heme binding /  metal ion binding metal ion bindingSimilarity search - Function | ||||||
| Biological species |   Elephas maximus (Asiatic elephant) Elephas maximus (Asiatic elephant) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.78 Å X-RAY DIFFRACTION / Resolution: 1.78 Å | ||||||
 Authors Authors | Bisig, D.A. / Piontek, K. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 1995 Journal: J.Biol.Chem. / Year: 1995Title: Crystal structure of Asian elephant (Elephas maximus) cyano-metmyoglobin at 1.78-A resolution. Phe29(B10) accounts for its unusual ligand binding properties. Authors: Bisig, D.A. / Di Iorio, E.E. / Diederichs, K. / Winterhalter, K.H. / Piontek, K. #1:  Journal: To be Published Journal: To be PublishedTitle: A Double Mutant of Sperm Whale Myoglobin Mimics the Structure and Function of Elephant Myoglobin Authors: Zhao, X. / Vyas, K. / Nguyen, B.D. / Rajarathnam, K. / La Mar, G.N. / Li, T. / Phillips Jr., G.N. / Eich, R.F. / Olson, J.S. / Ling, J. / Bocian, D.F. #2:  Journal: J.Biol.Chem. / Year: 1993 Journal: J.Biol.Chem. / Year: 1993Title: 1H NMR Investigation of the Heme Cavity of Elephant (E7 Gln) met-Cyano-Myoglobin. Evidence for a B-Helix Phenylalanine Interaction with Bound Ligand Authors: Vyas, K. / Rajarathnam, K. / Yu, L.P. / Emerson, S.D. / La Mar, G.N. / Krishnamoorthi, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1emy.cif.gz 1emy.cif.gz | 49.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1emy.ent.gz pdb1emy.ent.gz | 34.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1emy.json.gz 1emy.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/em/1emy https://data.pdbj.org/pub/pdb/validation_reports/em/1emy ftp://data.pdbj.org/pub/pdb/validation_reports/em/1emy ftp://data.pdbj.org/pub/pdb/validation_reports/em/1emy | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  Mass: 17026.592 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Elephas maximus (Asiatic elephant) / Organ: SKELETAL Elephas maximus (Asiatic elephant) / Organ: SKELETAL Skeleton / References: UniProt: P02186 Skeleton / References: UniProt: P02186 |
|---|---|
| #2: Chemical | ChemComp-CYN /  Cyanide Cyanide |
| #3: Chemical | ChemComp-HEM /  Heme B Heme B |
| #4: Water | ChemComp-HOH /  Water Water |
| Compound details | COMPND MET FORM. |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.03 Å3/Da / Density % sol: 39.41 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Details: CRYSTALLIZATION CONDITIONS METHOD: HANGING DROP VAPOR DIFFUSION PRECIPITANT: 45% PEG 1000 BUFFER: 0.1M TRIS/HCL PH: 8.5 SALT: 0.15M MAGNESIUM ACETATE PROTEIN CONCENTRATION: 10MG/ML ...Details: CRYSTALLIZATION CONDITIONS METHOD: HANGING DROP VAPOR DIFFUSION PRECIPITANT: 45% PEG 1000 BUFFER: 0.1M TRIS/HCL PH: 8.5 SALT: 0.15M MAGNESIUM ACETATE PROTEIN CONCENTRATION: 10MG/ML TEMPERATURE: 293K CRYSTAL SIZE: 0.8 X 0.4 X 0.3 MM | |||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 8.5 / Method: vapor diffusion, hanging drop / Details: macroseeding | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | Num. obs: 13649 / % possible obs: 93.6 % / Observed criterion σ(I): 0 |
| Reflection | *PLUS Highest resolution: 1.78 Å / Redundancy: 5 % / Num. measured all: 68768 / Rmerge(I) obs: 0.045 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.78→8 Å / σ(F): 2 /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 21.45 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.78→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller



















 PDBj
PDBj













