+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1dyo | ||||||
|---|---|---|---|---|---|---|---|
| Title | Xylan-Binding Domain from CBM 22, formally x6b domain | ||||||
 Components Components | ENDO-1,4-BETA-XYLANASE Y | ||||||
 Keywords Keywords |  CARBOHYDRATE-BINDING MODULE / XYLAN-BINDING / CARBOHYDRATE-BINDING MODULE / XYLAN-BINDING /  XYLANASE XYLANASE | ||||||
| Function / homology |  Function and homology information Function and homology information cellulosome / cellulosome /  endo-1,4-beta-xylanase activity / endo-1,4-beta-xylanase activity /  endo-1,4-beta-xylanase / xylan catabolic process endo-1,4-beta-xylanase / xylan catabolic processSimilarity search - Function | ||||||
| Biological species |   CLOSTRIDIUM THERMOCELLUM (bacteria) CLOSTRIDIUM THERMOCELLUM (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 2.1 Å MAD / Resolution: 2.1 Å | ||||||
 Authors Authors | Davies, G.J. / Charnock, S.J. / Gilbert, H.J. / Fontes, C.M.G.A. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2000 Journal: Biochemistry / Year: 2000Title: The X6 Thermostabilising Domains of Xylanases are Carbohydrate Binding Modules: Structure and Biochemistry of the Clostridium Thermocellum X6B Domain Authors: Charnock, S.J. / Bolam, D.N. / Turkenburg, J.P. / Gilbert, H.J. / Ferreira, L.M.A. / Davies, G.J. / Fontes, C.M.G.A. | ||||||
| History |
| ||||||
| Remark 700 | SHEET DETERMINATION METHOD: THERE ARE SEVERAL BIFURCATED SHEETS IN THIS STRUCTURE. EACH IS ... SHEET DETERMINATION METHOD: THERE ARE SEVERAL BIFURCATED SHEETS IN THIS STRUCTURE. EACH IS REPRESENTED BY TWO SHEETS WHICH HAVE ONE OR MORE IDENTICAL STRANDS. SHEETS A AND A1 REPRESENT ONE BIFURCATED SHEET IN CHAIN A SHEETS C AND C1 REPRESENT ONE BIFURCATED SHEET IN CHAIN B |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1dyo.cif.gz 1dyo.cif.gz | 77.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1dyo.ent.gz pdb1dyo.ent.gz | 62.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1dyo.json.gz 1dyo.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dy/1dyo https://data.pdbj.org/pub/pdb/validation_reports/dy/1dyo ftp://data.pdbj.org/pub/pdb/validation_reports/dy/1dyo ftp://data.pdbj.org/pub/pdb/validation_reports/dy/1dyo | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 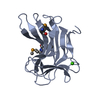
| ||||||||
| 2 | 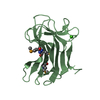
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.392, 0.815, 0.427), Vector  : : |
- Components
Components
| #1: Protein | Mass: 17862.248 Da / Num. of mol.: 2 / Fragment: XYLAN-BINDING DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)   CLOSTRIDIUM THERMOCELLUM (bacteria) / Production host: CLOSTRIDIUM THERMOCELLUM (bacteria) / Production host:   ESCHERICHIA COLI (E. coli) / References: UniProt: P51584 ESCHERICHIA COLI (E. coli) / References: UniProt: P51584#2: Chemical | #3: Water | ChemComp-HOH / |  Water WaterSequence details | RESIDUE ASN 565 FROM SWISSPROT ENTRY P51584 IS DELETED FROM THE PDB ENTRY | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.36 Å3/Da / Density % sol: 63.39 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | pH: 4.6 Details: CRYSTALS WERE GROWN FROM A PROTEIN CONCENTRATION OF 50 MG/ML IN 0.1 M NAAC BUFFER, PH 4.6, CONTAINING 10 MM DTT, 25 % (V/V) GLYCEROL AND WITH 12 % (W/V) PEG 8000 AS THE PRECIPITANT | ||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: BW7B / Wavelength: 0.8445 / Beamline: BW7B / Wavelength: 0.8445 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Oct 15, 1999 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.8445 Å / Relative weight: 1 : 0.8445 Å / Relative weight: 1 |
| Reflection | Resolution: 2.12→20 Å / Num. obs: 141380 / % possible obs: 98 % / Redundancy: 5.11 % / Rmerge(I) obs: 0.074 / Rsym value: 0.074 / Net I/σ(I): 19.2 |
| Reflection shell | Resolution: 2.12→2.2 Å / Redundancy: 4.6 % / Rmerge(I) obs: 0.437 / Mean I/σ(I) obs: 3.4 / Rsym value: 0.437 / % possible all: 96.5 |
| Reflection shell | *PLUS % possible obs: 96.5 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MAD / Resolution: 2.1→10 Å / Cross valid method: THROUGHOUT / σ(F): 0 MAD / Resolution: 2.1→10 Å / Cross valid method: THROUGHOUT / σ(F): 0 Details: FOR BOTH CHAINS THE FIRST 4 N- TERMINAL RESIDUES ARE DISORDERED
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj



