+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1c1l | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | LACTOSE-LIGANDED CONGERIN I | |||||||||
 Components Components | PROTEIN (CONGERIN I) | |||||||||
 Keywords Keywords | SUGAR BINDING PROTEIN /  GALECTIN / GALECTIN /  LECTIN / BETA-GALACTOSE-BINDING LECTIN / BETA-GALACTOSE-BINDING | |||||||||
| Function / homology |  Function and homology information Function and homology information galactoside binding / galactoside binding /  laminin binding / laminin binding /  carbohydrate binding / collagen-containing extracellular matrix / carbohydrate binding / collagen-containing extracellular matrix /  extracellular space extracellular spaceSimilarity search - Function | |||||||||
| Biological species |   Conger myriaster (whitespotted conger) Conger myriaster (whitespotted conger) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.5 Å MOLECULAR REPLACEMENT / Resolution: 1.5 Å | |||||||||
 Authors Authors | Shirai, T. / Mitsuyama, C. / Niwa, Y. / Matsui, Y. / Hotta, H. / Yamane, T. / Kamiya, H. / Ishii, C. / Ogawa, T. / Muramoto, K. | |||||||||
 Citation Citation |  Journal: Structure Fold.Des. / Year: 1999 Journal: Structure Fold.Des. / Year: 1999Title: High-resolution structure of the conger eel galectin, congerin I, in lactose-liganded and ligand-free forms: emergence of a new structure class by accelerated evolution. Authors: Shirai, T. / Mitsuyama, C. / Niwa, Y. / Matsui, Y. / Hotta, H. / Yamane, T. / Kamiya, H. / Ishii, C. / Ogawa, T. / Muramoto, K. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1c1l.cif.gz 1c1l.cif.gz | 43.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1c1l.ent.gz pdb1c1l.ent.gz | 28.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1c1l.json.gz 1c1l.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/c1/1c1l https://data.pdbj.org/pub/pdb/validation_reports/c1/1c1l ftp://data.pdbj.org/pub/pdb/validation_reports/c1/1c1l ftp://data.pdbj.org/pub/pdb/validation_reports/c1/1c1l | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1c1fC 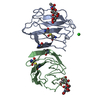 1sltS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 15356.081 Da / Num. of mol.: 1 / Fragment: CARBOHYDRATE-RECOGNITION-DOMAIN / Source method: isolated from a natural source / Details: COMPLEXED WITH LACTOSE / Source: (natural)   Conger myriaster (whitespotted conger) / Secretion: NON-CLASSICAL / Tissue: SKIN MUCUS / References: UniProt: P26788 Conger myriaster (whitespotted conger) / Secretion: NON-CLASSICAL / Tissue: SKIN MUCUS / References: UniProt: P26788 |
|---|---|
| #2: Polysaccharide | beta-D-galactopyranose-(1-4)-beta-D-glucopyranose / beta-lactose |
| #3: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.28 Å3/Da / Density % sol: 46 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | pH: 9 / Details: pH 9.0 | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 18 ℃ / pH: 7 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 291 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Photon Factory Photon Factory  / Beamline: BL-6A / Wavelength: 1 / Beamline: BL-6A / Wavelength: 1 |
| Detector | Type: FUJI / Detector: IMAGE PLATE |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.5→20 Å / Num. obs: 20509 / % possible obs: 89 % / Observed criterion σ(I): 2 / Redundancy: 2.2 % / Rmerge(I) obs: 0.068 |
| Reflection shell | Resolution: 1.5→1.55 Å / % possible all: 65.5 |
| Reflection shell | *PLUS % possible obs: 65.5 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1SLT Resolution: 1.5→8 Å / Cross valid method: THROUGHOUT / σ(F): 3 Details: SIDE-CHAINS OF GLN28, SER123 AND LEU124 ARE MODELED AS ALTERNATIVE CONFORMERS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.5→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS σ(F): 3 / % reflection Rfree: 5 % / Rfactor obs: 0.213 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj





