+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1by0 | ||||||
|---|---|---|---|---|---|---|---|
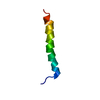
| Title | N-TERMINAL LEUCINE-REPEAT REGION OF HEPATITIS DELTA ANTIGEN | ||||||
 Components Components | PROTEIN (HEPATITIS DELTA ANTIGEN) | ||||||
 Keywords Keywords |  RNA BINDING PROTEIN / HEPATITIS DELTA ANTIGEN / RNA BINDING PROTEIN / HEPATITIS DELTA ANTIGEN /  HELIX / SOLUTION STRUCTURE / HELIX / SOLUTION STRUCTURE /  RNA BINDING RNA BINDING | ||||||
| Function / homology |  Function and homology information Function and homology information host cell / host cell /  virion component / viral penetration into host nucleus / symbiont entry into host cell / host cell nucleus / virion component / viral penetration into host nucleus / symbiont entry into host cell / host cell nucleus /  RNA binding RNA bindingSimilarity search - Function | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Cheng, J.W. / Lin, I.J. / Lou, Y.C. | ||||||
 Citation Citation |  Journal: Proteins / Year: 1999 Journal: Proteins / Year: 1999Title: Solution structure and RNA-binding activity of the N-terminal leucine-repeat region of hepatitis delta antigen Authors: Lin, I.J. / Lou, Y.C. / Pai, M.T. / Wu, H.N. / Cheng, J.W. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1by0.cif.gz 1by0.cif.gz | 227.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1by0.ent.gz pdb1by0.ent.gz | 193.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1by0.json.gz 1by0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/by/1by0 https://data.pdbj.org/pub/pdb/validation_reports/by/1by0 ftp://data.pdbj.org/pub/pdb/validation_reports/by/1by0 ftp://data.pdbj.org/pub/pdb/validation_reports/by/1by0 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 3475.154 Da / Num. of mol.: 1 / Fragment: RESIDUES 24-50 / Source method: obtained synthetically Details: THE PROTEIN WAS CHEMICALLY SYNTHESIZED FROM THE HEPATITIS D VIRUS References: UniProt: P25989 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: DISTANCE RESTRAINTS WERE REQUIRED FROM NOES ASSIGNED IN A 2D NOESY SPECTRUM WITH 75 MS MIXING TIME. |
- Sample preparation
Sample preparation
| Sample conditions | pH: 6.8 / Pressure: 1 atm / Temperature: 298 K |
|---|---|
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker DMX600 / Manufacturer: Bruker / Model : DMX600 / Field strength: 600 MHz : DMX600 / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 | ||||||||||||
| NMR ensemble | Conformer selection criteria: LOWEST TOTAL ENERGY / Conformers calculated total number: 200 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller










 PDBj
PDBj