+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1bhd | ||||||
|---|---|---|---|---|---|---|---|
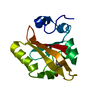
| Title | SECOND CALPONIN HOMOLOGY DOMAIN FROM UTROPHIN | ||||||
 Components Components | UTROPHIN | ||||||
 Keywords Keywords |  STRUCTURAL PROTEIN / CALPONIN HOMOLOGY / STRUCTURAL PROTEIN / CALPONIN HOMOLOGY /  ACTIN BINDING ACTIN BINDING | ||||||
| Function / homology |  Function and homology information Function and homology informationsynaptic signaling /  dystrophin-associated glycoprotein complex / regulation of sodium ion transmembrane transporter activity / dystrophin-associated glycoprotein complex / regulation of sodium ion transmembrane transporter activity /  vinculin binding / EGR2 and SOX10-mediated initiation of Schwann cell myelination / filopodium membrane / muscle organ development / positive regulation of cell-matrix adhesion / vinculin binding / EGR2 and SOX10-mediated initiation of Schwann cell myelination / filopodium membrane / muscle organ development / positive regulation of cell-matrix adhesion /  filopodium / filopodium /  muscle contraction ...synaptic signaling / muscle contraction ...synaptic signaling /  dystrophin-associated glycoprotein complex / regulation of sodium ion transmembrane transporter activity / dystrophin-associated glycoprotein complex / regulation of sodium ion transmembrane transporter activity /  vinculin binding / EGR2 and SOX10-mediated initiation of Schwann cell myelination / filopodium membrane / muscle organ development / positive regulation of cell-matrix adhesion / vinculin binding / EGR2 and SOX10-mediated initiation of Schwann cell myelination / filopodium membrane / muscle organ development / positive regulation of cell-matrix adhesion /  filopodium / filopodium /  muscle contraction / muscle contraction /  neuromuscular junction / neuromuscular junction /  sarcolemma / sarcolemma /  integrin binding / integrin binding /  actin binding / actin binding /  postsynaptic membrane / postsynaptic membrane /  cytoskeleton / cytoskeleton /  protein kinase binding / protein-containing complex / extracellular exosome / zinc ion binding / protein kinase binding / protein-containing complex / extracellular exosome / zinc ion binding /  nucleoplasm / nucleoplasm /  membrane / membrane /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Keep, N.H. / Winder, S.J. / Kendrick-Jones, J. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1999 Journal: J.Mol.Biol. / Year: 1999Title: The 2.0 A structure of the second calponin homology domain from the actin-binding region of the dystrophin homologue utrophin. Authors: Keep, N.H. / Norwood, F.L. / Moores, C.A. / Winder, S.J. / Kendrick-Jones, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1bhd.cif.gz 1bhd.cif.gz | 62 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1bhd.ent.gz pdb1bhd.ent.gz | 44 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1bhd.json.gz 1bhd.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bh/1bhd https://data.pdbj.org/pub/pdb/validation_reports/bh/1bhd ftp://data.pdbj.org/pub/pdb/validation_reports/bh/1bhd ftp://data.pdbj.org/pub/pdb/validation_reports/bh/1bhd | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1aa2S S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.601644, -0.050091, -0.797192), Vector  : : |
- Components
Components
| #1: Protein |  Mass: 13678.681 Da / Num. of mol.: 2 Fragment: 2ND CALPONIN HOMOLOGY DOMAIN FROM ACTIN BINDING REGION Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Tissue: MUSCLE Homo sapiens (human) / Tissue: MUSCLE Skeletal muscle / Description: EXPRESSED IN ESCHERICHIA COLI / Cell line: BL21 / Cellular location: LINKS CYTOSKELETON TO PLASMA MEMBRANE / Gene: UTRN, DMDL / Organ: PLASMA / Plasmid: PSJW1 (T7) / Species (production host): Escherichia coli / Cellular location (production host): CYTOPLASM / Gene (production host): UTRN DMDL / Production host: Skeletal muscle / Description: EXPRESSED IN ESCHERICHIA COLI / Cell line: BL21 / Cellular location: LINKS CYTOSKELETON TO PLASMA MEMBRANE / Gene: UTRN, DMDL / Organ: PLASMA / Plasmid: PSJW1 (T7) / Species (production host): Escherichia coli / Cellular location (production host): CYTOPLASM / Gene (production host): UTRN DMDL / Production host:   Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21 (DE3) / References: UniProt: P46939 Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21 (DE3) / References: UniProt: P46939#2: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.2 Å3/Da / Density % sol: 44.24 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | pH: 7.6 / Details: pH 7.6 | |||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / Method: vapor diffusion | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SRS SRS  / Beamline: PX9.5 / Wavelength: 1 / Beamline: PX9.5 / Wavelength: 1 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Aug 25, 1997 |
| Radiation | Monochromator: SI(111) / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2→22 Å / Num. obs: 14514 / % possible obs: 99.5 % / Observed criterion σ(I): 0 / Redundancy: 2.8 % / Biso Wilson estimate: 23.85 Å2 / Rmerge(I) obs: 0.032 / Net I/σ(I): 10.8 |
| Reflection shell | Resolution: 1.99→2.1 Å / Redundancy: 2.8 % / Rmerge(I) obs: 0.125 / Mean I/σ(I) obs: 5.9 / % possible all: 98.9 |
| Reflection | *PLUS Num. obs: 16459 / Num. measured all: 46728 |
| Reflection shell | *PLUS % possible obs: 98.9 % / Num. unique obs: 2296 / Num. measured obs: 6400 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB 1AA2 Resolution: 2→20 Å / Cross valid method: THROUGHOUT / σ(F): 0 Details: 0.011 AROMATIC PLANAR DEVIATION VALUE ABOVE IS FOR PEPTIDE. ESTIMATED COORDINATE ERROR. ESD FROM SIGMAA (A) : 0.185 LOW RESOLUTION CUTOFF (A) : 20.0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 31.23 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati d res low obs: 20 Å / Luzzati sigma a obs: 0.18 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: REFMAC / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.185 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Highest resolution: 2 Å / Lowest resolution: 2.13 Å / Rfactor Rfree: 0.319 / Num. reflection Rfree: 132 / % reflection Rfree: 5 % / Rfactor obs: 0.188 |
 Movie
Movie Controller
Controller












 PDBj
PDBj




