[English] 日本語
 Yorodumi
Yorodumi- PDB-1ba9: THE SOLUTION STRUCTURE OF REDUCED MONOMERIC SUPEROXIDE DISMUTASE,... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ba9 | ||||||
|---|---|---|---|---|---|---|---|
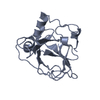
| Title | THE SOLUTION STRUCTURE OF REDUCED MONOMERIC SUPEROXIDE DISMUTASE, NMR, 36 STRUCTURES | ||||||
 Components Components | SUPEROXIDE DISMUTASE | ||||||
 Keywords Keywords |  OXIDOREDUCTASE / OXIDOREDUCTASE /  SUPEROXIDE DISMUTASE / COPPER-ZINC ENZYME / DISMUTATION OF UPEROXIDE RADICALS TO MOLECULAR OXYGEN AND HYDROGEN PEROXIDE SUPEROXIDE DISMUTASE / COPPER-ZINC ENZYME / DISMUTATION OF UPEROXIDE RADICALS TO MOLECULAR OXYGEN AND HYDROGEN PEROXIDE | ||||||
| Function / homology |  Function and homology information Function and homology informationaction potential initiation / neurofilament cytoskeleton organization / anterograde axonal transport / retrograde axonal transport / positive regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / protein phosphatase 2B binding / regulation of organ growth / relaxation of vascular associated smooth muscle / response to superoxide /  peripheral nervous system myelin maintenance ...action potential initiation / neurofilament cytoskeleton organization / anterograde axonal transport / retrograde axonal transport / positive regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / protein phosphatase 2B binding / regulation of organ growth / relaxation of vascular associated smooth muscle / response to superoxide / peripheral nervous system myelin maintenance ...action potential initiation / neurofilament cytoskeleton organization / anterograde axonal transport / retrograde axonal transport / positive regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / protein phosphatase 2B binding / regulation of organ growth / relaxation of vascular associated smooth muscle / response to superoxide /  peripheral nervous system myelin maintenance / regulation of T cell differentiation in thymus / superoxide anion generation / retina homeostasis / negative regulation of cholesterol biosynthetic process / hydrogen peroxide biosynthetic process / auditory receptor cell stereocilium organization / peripheral nervous system myelin maintenance / regulation of T cell differentiation in thymus / superoxide anion generation / retina homeostasis / negative regulation of cholesterol biosynthetic process / hydrogen peroxide biosynthetic process / auditory receptor cell stereocilium organization /  regulation of protein kinase activity / myeloid cell homeostasis / muscle cell cellular homeostasis / regulation of protein kinase activity / myeloid cell homeostasis / muscle cell cellular homeostasis /  regulation of GTPase activity / superoxide metabolic process / regulation of GTPase activity / superoxide metabolic process /  heart contraction / positive regulation of catalytic activity / heart contraction / positive regulation of catalytic activity /  superoxide dismutase / Detoxification of Reactive Oxygen Species / transmission of nerve impulse / negative regulation of reproductive process / negative regulation of developmental process / superoxide dismutase / Detoxification of Reactive Oxygen Species / transmission of nerve impulse / negative regulation of reproductive process / negative regulation of developmental process /  superoxide dismutase activity / neuronal action potential / regulation of multicellular organism growth / response to axon injury / ectopic germ cell programmed cell death / glutathione metabolic process / positive regulation of phagocytosis / ovarian follicle development / axon cytoplasm / superoxide dismutase activity / neuronal action potential / regulation of multicellular organism growth / response to axon injury / ectopic germ cell programmed cell death / glutathione metabolic process / positive regulation of phagocytosis / ovarian follicle development / axon cytoplasm /  embryo implantation / reactive oxygen species metabolic process / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / dendrite cytoplasm / removal of superoxide radicals / positive regulation of superoxide anion generation / regulation of mitochondrial membrane potential / thymus development / locomotory behavior / placenta development / response to organic substance / determination of adult lifespan / positive regulation of cytokine production / sensory perception of sound / response to hydrogen peroxide / embryo implantation / reactive oxygen species metabolic process / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / dendrite cytoplasm / removal of superoxide radicals / positive regulation of superoxide anion generation / regulation of mitochondrial membrane potential / thymus development / locomotory behavior / placenta development / response to organic substance / determination of adult lifespan / positive regulation of cytokine production / sensory perception of sound / response to hydrogen peroxide /  mitochondrial intermembrane space / mitochondrial intermembrane space /  small GTPase binding / small GTPase binding /  regulation of blood pressure / negative regulation of inflammatory response / regulation of blood pressure / negative regulation of inflammatory response /  peroxisome / Platelet degranulation / peroxisome / Platelet degranulation /  gene expression / response to heat / protein-folding chaperone binding / cytoplasmic vesicle / gene expression / response to heat / protein-folding chaperone binding / cytoplasmic vesicle /  spermatogenesis / response to ethanol / intracellular iron ion homeostasis / negative regulation of neuron apoptotic process / positive regulation of MAPK cascade / spermatogenesis / response to ethanol / intracellular iron ion homeostasis / negative regulation of neuron apoptotic process / positive regulation of MAPK cascade /  mitochondrial matrix / response to xenobiotic stimulus / positive regulation of apoptotic process / copper ion binding / neuronal cell body / apoptotic process / protein-containing complex / mitochondrial matrix / response to xenobiotic stimulus / positive regulation of apoptotic process / copper ion binding / neuronal cell body / apoptotic process / protein-containing complex /  mitochondrion / mitochondrion /  extracellular space / extracellular exosome / zinc ion binding / extracellular region / extracellular space / extracellular exosome / zinc ion binding / extracellular region /  nucleoplasm / identical protein binding / nucleoplasm / identical protein binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / TORTION ANGLES DYNAMIC (DYANA) SOLUTION NMR / TORTION ANGLES DYNAMIC (DYANA) | ||||||
 Authors Authors | Banci, L. / Benedetto, M. / Bertini, I. / Del Conte, R. / Piccioli, M. / Viezzoli, M.S. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1998 Journal: Biochemistry / Year: 1998Title: Solution structure of reduced monomeric Q133M2 copper, zinc superoxide dismutase (SOD). Why is SOD a dimeric enzyme?. Authors: Banci, L. / Benedetto, M. / Bertini, I. / Del Conte, R. / Piccioli, M. / Viezzoli, M.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ba9.cif.gz 1ba9.cif.gz | 1.5 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ba9.ent.gz pdb1ba9.ent.gz | 1.3 MB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ba9.json.gz 1ba9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ba/1ba9 https://data.pdbj.org/pub/pdb/validation_reports/ba/1ba9 ftp://data.pdbj.org/pub/pdb/validation_reports/ba/1ba9 ftp://data.pdbj.org/pub/pdb/validation_reports/ba/1ba9 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein |  / SOD / SODMass: 15832.447 Da / Num. of mol.: 1 / Mutation: C6A, F50E, G51E, C111S, E133Q Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: HSOD / Cellular location (production host): PERIPLASM / Gene (production host): HSOD / Production host: Homo sapiens (human) / Gene: HSOD / Cellular location (production host): PERIPLASM / Gene (production host): HSOD / Production host:   Escherichia coli (E. coli) / Strain (production host): TOPP 1 (STRATAGENE) / References: UniProt: P00441, Escherichia coli (E. coli) / Strain (production host): TOPP 1 (STRATAGENE) / References: UniProt: P00441,  superoxide dismutase superoxide dismutase |
|---|---|
| #2: Chemical | ChemComp-ZN / |
| #3: Chemical | ChemComp-CU1 /  Copper Copper |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NMR details | Text: THE STRUCTURE WAS DETERMINED USING TRIPLE-RESONANCE NMR SPECTROSCOPY ON 13C, 15N LABELED SOD. |
- Sample preparation
Sample preparation
| Details | Contents: H2O AND D2O |
|---|---|
| Sample conditions | pH: 5 / Pressure: 1 atm / Temperature: 298 K |
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| Software | Name:  AMBER / Classification: refinement AMBER / Classification: refinement | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR software |
| ||||||||||||||||
| Refinement | Method: TORTION ANGLES DYNAMIC (DYANA) / Software ordinal: 1 Details: REFINEMENT DETAILS CAN BE FOUND IN THE JRNL CITATION ABOVE. | ||||||||||||||||
| NMR ensemble | Conformer selection criteria: target function / Conformers calculated total number: 36 / Conformers submitted total number: 36 |
 Movie
Movie Controller
Controller











 PDBj
PDBj










