[English] 日本語
 Yorodumi
Yorodumi- PDB-1adl: ADIPOCYTE LIPID BINDING PROTEIN COMPLEXED WITH ARACHIDONIC ACID: ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1adl | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | ADIPOCYTE LIPID BINDING PROTEIN COMPLEXED WITH ARACHIDONIC ACID: X-RAY CRYSTALLOGRAPHIC AND TITRATION CALORIMETRY STUDIES | |||||||||
 Components Components | ADIPOCYTE LIPID-BINDING PROTEIN | |||||||||
 Keywords Keywords | LIPID BINDING PROTEIN / LIPID-BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationTriglyceride catabolism /  hormone receptor binding / long-chain fatty acid transmembrane transporter activity / cellular response to lithium ion / hormone receptor binding / long-chain fatty acid transmembrane transporter activity / cellular response to lithium ion /  long-chain fatty acid binding / white fat cell differentiation / fatty acid transport / long-chain fatty acid transport / brown fat cell differentiation / fatty acid metabolic process ...Triglyceride catabolism / long-chain fatty acid binding / white fat cell differentiation / fatty acid transport / long-chain fatty acid transport / brown fat cell differentiation / fatty acid metabolic process ...Triglyceride catabolism /  hormone receptor binding / long-chain fatty acid transmembrane transporter activity / cellular response to lithium ion / hormone receptor binding / long-chain fatty acid transmembrane transporter activity / cellular response to lithium ion /  long-chain fatty acid binding / white fat cell differentiation / fatty acid transport / long-chain fatty acid transport / brown fat cell differentiation / fatty acid metabolic process / cholesterol homeostasis / long-chain fatty acid binding / white fat cell differentiation / fatty acid transport / long-chain fatty acid transport / brown fat cell differentiation / fatty acid metabolic process / cholesterol homeostasis /  fatty acid binding / response to bacterium / positive regulation of inflammatory response / cellular response to tumor necrosis factor / positive regulation of cold-induced thermogenesis / negative regulation of DNA-templated transcription / positive regulation of cell population proliferation / fatty acid binding / response to bacterium / positive regulation of inflammatory response / cellular response to tumor necrosis factor / positive regulation of cold-induced thermogenesis / negative regulation of DNA-templated transcription / positive regulation of cell population proliferation /  nucleoplasm / nucleoplasm /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | |||||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.6 Å X-RAY DIFFRACTION / Resolution: 1.6 Å | |||||||||
 Authors Authors | Lalonde, J.M. / Levenson, M. / Roe, J.J. / Bernlohr, D.A. / Banaszak, L.J. | |||||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 1994 Journal: J.Biol.Chem. / Year: 1994Title: Adipocyte lipid-binding protein complexed with arachidonic acid. Titration calorimetry and X-ray crystallographic studies. Authors: LaLonde, J.M. / Levenson, M.A. / Roe, J.J. / Bernlohr, D.A. / Banaszak, L.J. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1adl.cif.gz 1adl.cif.gz | 41.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1adl.ent.gz pdb1adl.ent.gz | 27.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1adl.json.gz 1adl.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ad/1adl https://data.pdbj.org/pub/pdb/validation_reports/ad/1adl ftp://data.pdbj.org/pub/pdb/validation_reports/ad/1adl ftp://data.pdbj.org/pub/pdb/validation_reports/ad/1adl | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14587.688 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / References: UniProt: P04117 Mus musculus (house mouse) / References: UniProt: P04117 |
|---|---|
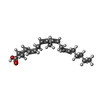
| #2: Chemical | ChemComp-ACD /  Arachidonic acid Arachidonic acid |
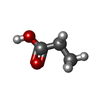
| #3: Chemical | ChemComp-PPI /  Propionic acid Propionic acid |
| #4: Water | ChemComp-HOH /  Water Water |
| Compound details | CYS 117 IS PARTIALLY OXIDIZED. |
| Nonpolymer details | AN UNKNOWN COMPOUND MODELED AS PROPANOIC ACID (PPI) WAS FOUND IN THE ELECTRON DENSITY MAP. |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.23 Å3/Da / Density % sol: 44.85 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | *PLUS Method: vapor diffusion, hanging drop / PH range low: 7.2 / PH range high: 6.6 | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | *PLUS Highest resolution: 1.6 Å / Num. obs: 16949 / % possible obs: 96 % / Observed criterion σ(I): 2 / Num. measured all: 70836 / Rmerge(I) obs: 0.04 |
| Reflection shell | *PLUS Mean I/σ(I) obs: 5.7 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.6→10 Å / σ(F): 1
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.188 / Rfactor Rfree : 0.22 : 0.22 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 16.6 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: x_angle_d / Dev ideal: 1.39 |
 Movie
Movie Controller
Controller












 PDBj
PDBj