+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8242 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | EBOV GP in complex with IgG c4G7 and c13C6 Fabs | ||||||||||||
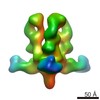
 Map data Map data | EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6 | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationhost cell endoplasmic reticulum / viral budding from plasma membrane / clathrin-dependent endocytosis of virus by host cell / suppression by virus of host tetherin activity / host cell cytoplasm / entry receptor-mediated virion attachment to host cell / symbiont-mediated suppression of host innate immune response / symbiont entry into host cell / membrane raft / fusion of virus membrane with host endosome membrane ...host cell endoplasmic reticulum / viral budding from plasma membrane / clathrin-dependent endocytosis of virus by host cell / suppression by virus of host tetherin activity / host cell cytoplasm / entry receptor-mediated virion attachment to host cell / symbiont-mediated suppression of host innate immune response / symbiont entry into host cell / membrane raft / fusion of virus membrane with host endosome membrane / lipid binding / viral envelope / host cell plasma membrane / virion membrane / extracellular region / identical protein binding Similarity search - Function | ||||||||||||
| Biological species |    Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.3 Å | ||||||||||||
 Authors Authors | Pallesen J / Murin CD / de Val N / Cottrell CA / Hastie KM / Turner HL / Fusco ML / Flyak AI / Zeitlin L / Crowe Jr JE ...Pallesen J / Murin CD / de Val N / Cottrell CA / Hastie KM / Turner HL / Fusco ML / Flyak AI / Zeitlin L / Crowe Jr JE / Andersen KG / Saphire EO / Ward AB | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2016 Journal: Nat Microbiol / Year: 2016Title: Structures of Ebola virus GP and sGP in complex with therapeutic antibodies. Authors: Jesper Pallesen / Charles D Murin / Natalia de Val / Christopher A Cottrell / Kathryn M Hastie / Hannah L Turner / Marnie L Fusco / Andrew I Flyak / Larry Zeitlin / James E Crowe / Kristian ...Authors: Jesper Pallesen / Charles D Murin / Natalia de Val / Christopher A Cottrell / Kathryn M Hastie / Hannah L Turner / Marnie L Fusco / Andrew I Flyak / Larry Zeitlin / James E Crowe / Kristian G Andersen / Erica Ollmann Saphire / Andrew B Ward /  Abstract: The Ebola virus (EBOV) GP gene encodes two glycoproteins. The major product is a soluble, dimeric glycoprotein (sGP) that is secreted abundantly. Despite the abundance of sGP during infection, little ...The Ebola virus (EBOV) GP gene encodes two glycoproteins. The major product is a soluble, dimeric glycoprotein (sGP) that is secreted abundantly. Despite the abundance of sGP during infection, little is known regarding its structure or functional role. A minor product, resulting from transcriptional editing, is the transmembrane-anchored, trimeric viral surface glycoprotein (GP). GP mediates attachment to and entry into host cells, and is the intended target of antibody therapeutics. Because large portions of sequence are shared between GP and sGP, it has been hypothesized that sGP may potentially subvert the immune response or may contribute to pathogenicity. In this study, we present cryo-electron microscopy structures of GP and sGP in complex with GP-specific and GP/sGP cross-reactive antibodies undergoing human clinical trials. The structure of the sGP dimer presented here, in complex with both an sGP-specific antibody and a GP/sGP cross-reactive antibody, permits us to unambiguously assign the oligomeric arrangement of sGP and compare its structure and epitope presentation to those of GP. We also provide biophysical evaluation of naturally occurring GP/sGP mutations that fall within the footprints identified by our high-resolution structures. Taken together, our data provide a detailed and more complete picture of the accessible Ebolavirus glycoprotein landscape and a structural basis to evaluate patient and vaccine antibody responses towards differently structured products of the GP gene. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8242.map.gz emd_8242.map.gz | 39.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8242-v30.xml emd-8242-v30.xml emd-8242.xml emd-8242.xml | 32.1 KB 32.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8242_fsc_1.xml emd_8242_fsc_1.xml emd_8242_fsc_2.xml emd_8242_fsc_2.xml | 8.4 KB 8.5 KB | Display Display |  FSC data file FSC data file |
| Images |  emd_8242.png emd_8242.png | 121.6 KB | ||
| Others |  emd_8242_additional_1.map.gz emd_8242_additional_1.map.gz emd_8242_additional_2.map.gz emd_8242_additional_2.map.gz emd_8242_additional_3.map.gz emd_8242_additional_3.map.gz emd_8242_additional_4.map.gz emd_8242_additional_4.map.gz emd_8242_additional_5.map.gz emd_8242_additional_5.map.gz emd_8242_half_map_1.map.gz emd_8242_half_map_1.map.gz emd_8242_half_map_2.map.gz emd_8242_half_map_2.map.gz | 50.9 MB 51.6 MB 43 MB 43 MB 50.9 MB 43 MB 42.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8242 http://ftp.pdbj.org/pub/emdb/structures/EMD-8242 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8242 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8242 | HTTPS FTP |
-Validation report
| Summary document |  emd_8242_validation.pdf.gz emd_8242_validation.pdf.gz | 427.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8242_full_validation.pdf.gz emd_8242_full_validation.pdf.gz | 427.4 KB | Display | |
| Data in XML |  emd_8242_validation.xml.gz emd_8242_validation.xml.gz | 6 KB | Display | |
| Data in CIF |  emd_8242_validation.cif.gz emd_8242_validation.cif.gz | 6.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8242 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8242 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8242 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8242 | HTTPS FTP |
-Related structure data
| Related structure data |  5kenMC  8240C  8241C  5kelC  5kemC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8242.map.gz / Format: CCP4 / Size: 55.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8242.map.gz / Format: CCP4 / Size: 55.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: combined map; unfiltered and unsharpened
| File | emd_8242_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | combined map; unfiltered and unsharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6
| File | emd_8242_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6
| File | emd_8242_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6
| File | emd_8242_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6
| File | emd_8242_additional_5.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EBOV GP in complex with variable Fab domains of IgGs c4G7 and c13C6 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Supplemental map: emd 8242 half map 1.map
| File | emd_8242_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Supplemental map: emd 8242 half map 2.map
| File | emd_8242_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ebola virus trimeric surface glycoprotein in complex with IgG c4G...
| Entire | Name: Ebola virus trimeric surface glycoprotein in complex with IgG c4G7 and c13C6 Fabs |
|---|---|
| Components |
|
-Supramolecule #1: Ebola virus trimeric surface glycoprotein in complex with IgG c4G...
| Supramolecule | Name: Ebola virus trimeric surface glycoprotein in complex with IgG c4G7 and c13C6 Fabs type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Molecular weight | Experimental: 540 KDa |
-Macromolecule #1: Ebola surface glycoprotein, GP1
| Macromolecule | Name: Ebola surface glycoprotein, GP1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 30.910719 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: IPLGVIHNST LQVSDVDKLV CRDKLSSTNQ LRSVGLNLEG NGVATDVPSA TKRWGFRSGV PPKVVNYEAG EWAENCYNLE IKKPDGSEC LPAAPDGIRG FPRCRYVHKV SGTGPCAGDF AFHKEGAFFL YDRLASTVIY RGTTFAEGVV AFLILPQAKK D FFSSHPLR ...String: IPLGVIHNST LQVSDVDKLV CRDKLSSTNQ LRSVGLNLEG NGVATDVPSA TKRWGFRSGV PPKVVNYEAG EWAENCYNLE IKKPDGSEC LPAAPDGIRG FPRCRYVHKV SGTGPCAGDF AFHKEGAFFL YDRLASTVIY RGTTFAEGVV AFLILPQAKK D FFSSHPLR EPVNATEDPS SGYYSTTIRY QATGFGTNET EYLFEVDNLT YVQLESRFTP QFLLQLNETI YTSGKRSNTT GK LIWKVNP EIDTTIGEWA FWETKKNLTR KIRSEELSF |
-Macromolecule #2: Ebola surface glycoprotein, GP2
| Macromolecule | Name: Ebola surface glycoprotein, GP2 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.650368 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AIVNAQPKCN PNLHYWTTQD EGAAIGLAWI PYFGPAAEGI YTEGLMHNQD GLICGLRQLA NETTQALQLF LRATTELRTF SILNRKAID FLLQRWGGTC HILGPDCCIE PHDW |
-Macromolecule #3: c4G7 variable Fab domain heavy chain
| Macromolecule | Name: c4G7 variable Fab domain heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.85123 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EVQLQESGPE LEMPGASVKI SCKASGSSFT GFSMNWVKQS NGKSLEWIGN IDTYYGGTTY NQKFKGKATL TVDKSSSTAY MQLKSLTSE DSAVYYCARS AYYGSTFAYW GQGTLVTVS |
-Macromolecule #4: c4G7 variable Fab domain light chain
| Macromolecule | Name: c4G7 variable Fab domain light chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.733013 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DIQMTQSPAS LSASVGETVT ITCRASENIY SYLAWYQQKQ GKSPQLLVYN AKTLIEGVPS RFSGSGSGTQ FSLKINSLQP EDFGSYFCQ HHFGTPFTFG SGTELEIK |
-Macromolecule #5: c13C6 variable Fab domain light chain
| Macromolecule | Name: c13C6 variable Fab domain light chain / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.611877 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DIVMTQSPLS LSTSVGDRVS LTCKASQNVG TAVAWYQQKP GQSPKLLIYS ASNRYTGVPD RFTGSGSGTD FTLTISNMQS EDLADYFCQ QYSSYPLTFG AGTKLELR |
-Macromolecule #6: c13C6 variable Fab domain heavy chain
| Macromolecule | Name: c13C6 variable Fab domain heavy chain / type: protein_or_peptide / ID: 6 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.169679 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVKLLESGGG LVQPGGSLKL SCAASGFSLS TSGVGVGWFR QPSGKGLEWL ALIWWDDDKY YNPSLKSQLS ISKDFSRNQV FLKISNVDI ADTATYYCAR RDPFGYDNAM GYWGQGTSVT VS |
-Macromolecule #9: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 9 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Name: 1xTBS |
| Grid | Model: C-flat-2/2 4C / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 20.0 nm / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: OTHER / Pretreatment - Pressure: 0.0093 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 277 K / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77.0 K / Max: 77.0 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-50 / Number grids imaged: 1 / Number real images: 2555 / Average exposure time: 10.0 sec. / Average electron dose: 57.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 22500 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Model building and refinement were conducted using a combination of software programs. The refinement target was optimizing using the MolProbity score while maintaining a high (good) EMRinger score. |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: EMRinger |
| Output model |  PDB-5ken: |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X



























































