[English] 日本語
 Yorodumi
Yorodumi- EMDB-43683: Cryo-EM structure of FLVCR2 in the inward-facing state with choli... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
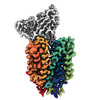
| Title | Cryo-EM structure of FLVCR2 in the inward-facing state with choline bound | |||||||||
 Map data Map data | FLVCR2 in the inward-facing state in complex with choline and Fab FLV23 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Choline transporter / Choline transporter /  blood-brain barrier / blood-brain barrier /  membrane protein / MFS fold / membrane protein / MFS fold /  TRANSPORT PROTEIN TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationheme transmembrane transporter activity /  mitochondrial membrane / mitochondrial membrane /  heme binding / endoplasmic reticulum membrane heme binding / endoplasmic reticulum membraneSimilarity search - Function | |||||||||
| Biological species |   Mus musculus (house mouse) / synthetic construct (others) Mus musculus (house mouse) / synthetic construct (others) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.77 Å cryo EM / Resolution: 2.77 Å | |||||||||
 Authors Authors | Cater RJ / Mancia F | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Structural and molecular basis of choline uptake into the brain by FLVCR2. Authors: Rosemary J Cater / Dibyanti Mukherjee / Eva Gil-Iturbe / Satchal K Erramilli / Ting Chen / Katie Koo / Nicolás Santander / Andrew Reckers / Brian Kloss / Tomasz Gawda / Brendon C Choy / ...Authors: Rosemary J Cater / Dibyanti Mukherjee / Eva Gil-Iturbe / Satchal K Erramilli / Ting Chen / Katie Koo / Nicolás Santander / Andrew Reckers / Brian Kloss / Tomasz Gawda / Brendon C Choy / Zhening Zhang / Aditya Katewa / Amara Larpthaveesarp / Eric J Huang / Scott W J Mooney / Oliver B Clarke / Sook Wah Yee / Kathleen M Giacomini / Anthony A Kossiakoff / Matthias Quick / Thomas Arnold / Filippo Mancia /    Abstract: Choline is an essential nutrient that the human body needs in vast quantities for cell membrane synthesis, epigenetic modification and neurotransmission. The brain has a particularly high demand for ...Choline is an essential nutrient that the human body needs in vast quantities for cell membrane synthesis, epigenetic modification and neurotransmission. The brain has a particularly high demand for choline, but how it enters the brain remains unknown. The major facilitator superfamily transporter FLVCR1 (also known as MFSD7B or SLC49A1) was recently determined to be a choline transporter but is not highly expressed at the blood-brain barrier, whereas the related protein FLVCR2 (also known as MFSD7C or SLC49A2) is expressed in endothelial cells at the blood-brain barrier. Previous studies have shown that mutations in human Flvcr2 cause cerebral vascular abnormalities, hydrocephalus and embryonic lethality, but the physiological role of FLVCR2 is unknown. Here we demonstrate both in vivo and in vitro that FLVCR2 is a BBB choline transporter and is responsible for the majority of choline uptake into the brain. We also determine the structures of choline-bound FLVCR2 in both inward-facing and outward-facing states using cryo-electron microscopy. These results reveal how the brain obtains choline and provide molecular-level insights into how FLVCR2 binds choline in an aromatic cage and mediates its uptake. Our work could provide a novel framework for the targeted delivery of therapeutic agents into the brain. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43683.map.gz emd_43683.map.gz | 197.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43683-v30.xml emd-43683-v30.xml emd-43683.xml emd-43683.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_43683.png emd_43683.png | 113.5 KB | ||
| Filedesc metadata |  emd-43683.cif.gz emd-43683.cif.gz | 6.4 KB | ||
| Others |  emd_43683_half_map_1.map.gz emd_43683_half_map_1.map.gz emd_43683_half_map_2.map.gz emd_43683_half_map_2.map.gz | 194.1 MB 194.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43683 http://ftp.pdbj.org/pub/emdb/structures/EMD-43683 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43683 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43683 | HTTPS FTP |
-Related structure data
| Related structure data |  8vznMC  8vzoC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43683.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43683.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | FLVCR2 in the inward-facing state in complex with choline and Fab FLV23 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: FLVCR2 in the inward-facing state in complex with...
| File | emd_43683_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | FLVCR2 in the inward-facing state in complex with choline and Fab FLV9 half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: FLVCR2 in the inward-facing state in complex with...
| File | emd_43683_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | FLVCR2 in the inward-facing state in complex with choline and Fab FLV9 half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : FLVCR2 in the inward-facing state complexed with choline and Fab FLV9
| Entire | Name: FLVCR2 in the inward-facing state complexed with choline and Fab FLV9 |
|---|---|
| Components |
|
-Supramolecule #1: FLVCR2 in the inward-facing state complexed with choline and Fab FLV9
| Supramolecule | Name: FLVCR2 in the inward-facing state complexed with choline and Fab FLV9 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
| Molecular weight | Theoretical: 110 KDa |
-Macromolecule #1: Feline leukemia virus subgroup C cellular receptor 2
| Macromolecule | Name: Feline leukemia virus subgroup C cellular receptor 2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
| Molecular weight | Theoretical: 60.103266 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MVNESLNQEE SNDRPAPESE FQMDTSYSTQ PSGSIHPSVS GHPSVSGHPS VSGHPSVSIH PSVSIDPSVS VRPSSSALPS TLAQPSGLT HHSSLVREDS VIKVSKRRWV VVLVFSCYSL CNAFQWIQYG SINNIFMNFY GVSAFAIDWL SMCYMLTYIP L LLPVAWML ...String: MVNESLNQEE SNDRPAPESE FQMDTSYSTQ PSGSIHPSVS GHPSVSGHPS VSGHPSVSIH PSVSIDPSVS VRPSSSALPS TLAQPSGLT HHSSLVREDS VIKVSKRRWV VVLVFSCYSL CNAFQWIQYG SINNIFMNFY GVSAFAIDWL SMCYMLTYIP L LLPVAWML EKFGLRTIAI TGSALNCLGA WVKLGSLEPH LFPVTMVGQV ICSVAQVFIL GMPSRIASVW FGADEVSTAC SV AVFGNQL GIAIGFLVPP VLVPNIKDPE KLAYHISIMF YIIGGVATFL FILVIIVFKE KPKHPPSRAQ SLSYALATTD ASY LSSIVR LFKNLNFVLL VITYGLNAGA FYALSTLLNR MVILHFPGEE VNAGRIGLTI VIAGMFGAMI SGIWLDKSKT YKET TLVVY IMTLVGMVVY TFTLNLNHLW VVFITAGTLG FFMTGYLPLG FEFAVELTYP ESEGVSSGLL NVSAQVFGIV FTISQ GQII DNHGTMFGNI FLCVFLALGS ALTAFIKSDL RRQRANKDAP ETKVQEEEEE EEGSNTSKVP VVSEAHL UniProtKB: Feline leukemia virus subgroup C cellular receptor 2 |
-Macromolecule #2: Fab FLV9 heavy chain
| Macromolecule | Name: Fab FLV9 heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 25.767727 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NVSSSYIHWV RQAPGKGLEW VASIYPYSGY TSYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARKQYNWPWP YIHSRYWGFD YWGQGTLVTV SSASTKGPSV FPLAPSSKST SGGTAALGCL V KDYFPEPV ...String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NVSSSYIHWV RQAPGKGLEW VASIYPYSGY TSYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARKQYNWPWP YIHSRYWGFD YWGQGTLVTV SSASTKGPSV FPLAPSSKST SGGTAALGCL V KDYFPEPV TVSWNSGALT SGVHTFPAVL QSSGLYSLSS VVTVPSSSLG TQTYICNVNH KPSNTKVDKK VEPKSCDKTH T |
-Macromolecule #3: Fab FLV9 light chain
| Macromolecule | Name: Fab FLV9 light chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 23.290828 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: SDIQMTQSPS SLSASVGDRV TITCRASQSV SSAVAWYQQK PGKAPKLLIY SASSLYSGVP SRFSGSRSGT DFTLTISSLQ PEDFATYYC QQSYGSLVTF GQGTKVEIKR TVAAPSVFIF PPSDSQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD ...String: SDIQMTQSPS SLSASVGDRV TITCRASQSV SSAVAWYQQK PGKAPKLLIY SASSLYSGVP SRFSGSRSGT DFTLTISSLQ PEDFATYYC QQSYGSLVTF GQGTKVEIKR TVAAPSVFIF PPSDSQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGEC |
-Macromolecule #4: CHOLINE ION
| Macromolecule | Name: CHOLINE ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: CHT |
|---|---|
| Molecular weight | Theoretical: 104.171 Da |
| Chemical component information |  ChemComp-CHT: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 58.2 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.77 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 308701 |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X