[English] 日本語
 Yorodumi
Yorodumi- EMDB-42451: Cryo-EM structure of a Clockwise locked form of the Salmonella en... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a Clockwise locked form of the Salmonella enterica Typhimurium flagellar C-ring, with C34 symmetry applied | |||||||||||||||
 Map data Map data | Clockwise locked form of the Salmonella enterica Typhimurium flagellar C-ring, with C34 symmetry applied | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords |  Flagella / Flagella /  Salmonella / Salmonella /  C-ring / C-ring /  motor / motor /  MOTOR PROTEIN MOTOR PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum basal body, MS ring / bacterial-type flagellum basal body / bacterial-type flagellum-dependent swarming motility / positive chemotaxis / cytoskeletal motor activity / bacterial-type flagellum-dependent cell motility /  chemotaxis / chemotaxis /  plasma membrane plasma membraneSimilarity search - Function | |||||||||||||||
| Biological species |   Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) | |||||||||||||||
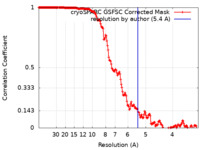
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 5.4 Å cryo EM / Resolution: 5.4 Å | |||||||||||||||
 Authors Authors | Johnson S / Deme JC / Lea SM | |||||||||||||||
| Funding support |  United States, United States,  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2024 Journal: Nat Microbiol / Year: 2024Title: Structural basis of directional switching by the bacterial flagellum. Authors: Steven Johnson / Justin C Deme / Emily J Furlong / Joseph J E Caesar / Fabienne F V Chevance / Kelly T Hughes / Susan M Lea /    Abstract: The bacterial flagellum is a macromolecular protein complex that harvests energy from uni-directional ion flow across the inner membrane to power bacterial swimming via rotation of the flagellar ...The bacterial flagellum is a macromolecular protein complex that harvests energy from uni-directional ion flow across the inner membrane to power bacterial swimming via rotation of the flagellar filament. Rotation is bi-directional, with binding of a cytoplasmic chemotactic response regulator controlling reversal, though the structural and mechanistic bases for rotational switching are not well understood. Here we present cryoelectron microscopy structures of intact Salmonella flagellar basal bodies (3.2-5.5 Å), including the cytoplasmic C-ring complexes required for power transmission, in both counter-clockwise and clockwise rotational conformations. These reveal 180° movements of both the N- and C-terminal domains of the FliG protein, which, when combined with a high-resolution cryoelectron microscopy structure of the MotAB stator, show that the stator shifts from the outside to the inside of the C-ring. This enables rotational switching and reveals how uni-directional ion flow across the inner membrane is used to accomplish bi-directional rotation of the flagellum. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42451.map.gz emd_42451.map.gz | 247.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42451-v30.xml emd-42451-v30.xml emd-42451.xml emd-42451.xml | 21 KB 21 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42451_fsc.xml emd_42451_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_42451.png emd_42451.png | 106.1 KB | ||
| Masks |  emd_42451_msk_1.map emd_42451_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-42451.cif.gz emd-42451.cif.gz | 6.4 KB | ||
| Others |  emd_42451_additional_1.map.gz emd_42451_additional_1.map.gz emd_42451_half_map_1.map.gz emd_42451_half_map_1.map.gz emd_42451_half_map_2.map.gz emd_42451_half_map_2.map.gz | 479.9 MB 474.2 MB 474.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42451 http://ftp.pdbj.org/pub/emdb/structures/EMD-42451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42451 | HTTPS FTP |
-Related structure data
| Related structure data |  8uplMC  8ucsC  8umdC  8umxC  8uoxC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_42451.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42451.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Clockwise locked form of the Salmonella enterica Typhimurium flagellar C-ring, with C34 symmetry applied | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.664 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_42451_msk_1.map emd_42451_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Sharpened map
| File | emd_42451_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map B
| File | emd_42451_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map A
| File | emd_42451_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Intact C34 flagellar C-ring from purified basal bodies, clockwise...
| Entire | Name: Intact C34 flagellar C-ring from purified basal bodies, clockwise locked. |
|---|---|
| Components |
|
-Supramolecule #1: Intact C34 flagellar C-ring from purified basal bodies, clockwise...
| Supramolecule | Name: Intact C34 flagellar C-ring from purified basal bodies, clockwise locked. type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
-Macromolecule #1: Flagellar M-ring protein
| Macromolecule | Name: Flagellar M-ring protein / type: protein_or_peptide / ID: 1 / Number of copies: 34 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 61.295645 KDa |
| Sequence | String: MSATASTATQ PKPLEWLNRL RANPRIPLIV AGSAAVAIVV AMVLWAKTPD YRTLFSNLSD QDGGAIVAQL TQMNIPYRFA NGSGAIEVP ADKVHELRLR LAQQGLPKGG AVGFELLDQE KFGISQFSEQ VNYQRALEGE LARTIETLGP VKSARVHLAM P KPSLFVRE ...String: MSATASTATQ PKPLEWLNRL RANPRIPLIV AGSAAVAIVV AMVLWAKTPD YRTLFSNLSD QDGGAIVAQL TQMNIPYRFA NGSGAIEVP ADKVHELRLR LAQQGLPKGG AVGFELLDQE KFGISQFSEQ VNYQRALEGE LARTIETLGP VKSARVHLAM P KPSLFVRE QKSPSASVTV TLEPGRALDE GQISAVVHLV SSAVAGLPPG NVTLVDQSGH LLTQSNTSGR DLNDAQLKFA ND VESRIQR RIEAILSPIV GNGNVHAQVT AQLDFANKEQ TEEHYSPNGD ASKATLRSRQ LNISEQVGAG YPGGVPGALS NQP APPNEA PIATPPTNQQ NAQNTPQTST STNSNSAGPR STQRNETSNY EVDRTIRHTK MNVGDIERLS VAVVVNYKTL ADGK PLPLT ADQMKQIEDL TREAMGFSDK RGDTLNVVNS PFSAVDNTGG ELPFWQQQSF IDQLLAAGRW LLVLVVAWIL WRKAV RPQL TRRVEEAKAA QEQAQVRQET EEAVEVRLSK DEQLQQRRAN QRLGAEVMSQ RIREMSDNDP RVVALVIRQW MSNDHE UniProtKB: Flagellar M-ring protein |
-Macromolecule #2: Flagellar motor switch protein FliG
| Macromolecule | Name: Flagellar motor switch protein FliG / type: protein_or_peptide / ID: 2 Details: Residues 169-171 (PAA) were deleted on the chromosome. Number of copies: 34 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 36.651688 KDa |
| Sequence | String: MSNLSGTDKS VILLMTIGED RAAEVFKHLS TREVQALSTA MANVRQISNK QLTDVLSEFE QEAEQFAALN INANEYLRSV LVKALGEER ASSLLEDILE TRDTTSGIET LNFMEPQSAA DLIRDEHPQI IATILVHLKR SQAADILALF DERLRHDVML R IATFGGVQ ...String: MSNLSGTDKS VILLMTIGED RAAEVFKHLS TREVQALSTA MANVRQISNK QLTDVLSEFE QEAEQFAALN INANEYLRSV LVKALGEER ASSLLEDILE TRDTTSGIET LNFMEPQSAA DLIRDEHPQI IATILVHLKR SQAADILALF DERLRHDVML R IATFGGVQ LAELTEVLNG LLDGQNLKRS KMGGVRTAAE IINLMKTQQE EAVITAVREF DGELAQKIID EMFLFENLVD VD DRSIQRL LQEVDSESLL IALKGAEPPL REKFLRNMSQ RAADILRDDL ANRGPVRLSQ VENEQKAILL IVRRLAETGE MVI GSGEDT YV UniProtKB:  Flagellar motor switch protein FliG Flagellar motor switch protein FliG |
-Macromolecule #3: Flagellar motor switch protein FliM
| Macromolecule | Name: Flagellar motor switch protein FliM / type: protein_or_peptide / ID: 3 / Number of copies: 34 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 37.901066 KDa |
| Sequence | String: MGDSILSQAE IDALLNGDSD TKDEPTPGIA SDSDIRPYDP NTQRRVVRER LQALEIINER FARQFRMGLF NLLRRSPDIT VGAIRIQPY HEFARNLPVP TNLNLIHLKP LRGTGLVVFS PSLVFIAVDN LFGGDGRFPT KVEGREFTHT EQRVINRMLK L ALEGYSDA ...String: MGDSILSQAE IDALLNGDSD TKDEPTPGIA SDSDIRPYDP NTQRRVVRER LQALEIINER FARQFRMGLF NLLRRSPDIT VGAIRIQPY HEFARNLPVP TNLNLIHLKP LRGTGLVVFS PSLVFIAVDN LFGGDGRFPT KVEGREFTHT EQRVINRMLK L ALEGYSDA WKAINPLEVE YVRSEMQVKF TNITTSPNDI VVNTPFHVEI GNLTGEFNIC LPFSMIEPLR ELLVNPPLEN SR HEDQNWR DNLVRQVQHS ELELVANFAD IPLRLSQILK LKPGDVLPIE KPDRIIAHVD GVPVLTSQYG TVNGQYALRV EHL INPILN SLNEEQPK UniProtKB:  Flagellar motor switch protein FliM Flagellar motor switch protein FliM |
-Macromolecule #4: Flagellar motor switch protein FliN
| Macromolecule | Name: Flagellar motor switch protein FliN / type: protein_or_peptide / ID: 4 / Number of copies: 102 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 14.801823 KDa |
| Sequence | String: MSDMNNPSDE NTGALDDLWA DALNEQKATT TKSAADAVFQ QLGGGDVSGA MQDIDLIMDI PVKLTVELGR TRMTIKELLR LTQGSVVAL DGLAGEPLDI LINGYLIAQG EVVVVADKYG VRITDIITPS ERMRRLSR UniProtKB:  Flagellar motor switch protein FliN Flagellar motor switch protein FliN |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 58.5 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z
Z Y
Y X
X