[English] 日本語
 Yorodumi
Yorodumi- EMDB-41840: Gaussian Mixture Models based single particle refinement - GPCR (... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Gaussian Mixture Models based single particle refinement - GPCR (Substance P bound to active human neurokinin 1 receptor in complex with miniGs399 from EMPIAR-10786) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  GPCR / GPCR /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsubstance P receptor binding / substance P receptor activity /  insemination / insemination /  tachykinin receptor activity / positive regulation of flagellated sperm motility / tachykinin receptor activity / positive regulation of flagellated sperm motility /  aggressive behavior / Tachykinin receptors bind tachykinins / positive regulation of uterine smooth muscle contraction / detection of abiotic stimulus / positive regulation of synaptic transmission, cholinergic ...substance P receptor binding / substance P receptor activity / aggressive behavior / Tachykinin receptors bind tachykinins / positive regulation of uterine smooth muscle contraction / detection of abiotic stimulus / positive regulation of synaptic transmission, cholinergic ...substance P receptor binding / substance P receptor activity /  insemination / insemination /  tachykinin receptor activity / positive regulation of flagellated sperm motility / tachykinin receptor activity / positive regulation of flagellated sperm motility /  aggressive behavior / Tachykinin receptors bind tachykinins / positive regulation of uterine smooth muscle contraction / detection of abiotic stimulus / positive regulation of synaptic transmission, cholinergic / aggressive behavior / Tachykinin receptors bind tachykinins / positive regulation of uterine smooth muscle contraction / detection of abiotic stimulus / positive regulation of synaptic transmission, cholinergic /  operant conditioning / operant conditioning /  sperm head / sperm ejaculation / positive regulation of lymphocyte proliferation / tachykinin receptor signaling pathway / response to ozone / response to auditory stimulus / positive regulation of action potential / smooth muscle contraction involved in micturition / positive regulation of blood pressure / regulation of smooth muscle cell proliferation / positive regulation of hormone secretion / positive regulation of vascular permeability / positive regulation of ossification / regulation of smooth muscle cell migration / positive regulation of leukocyte migration / response to pain / sperm head / sperm ejaculation / positive regulation of lymphocyte proliferation / tachykinin receptor signaling pathway / response to ozone / response to auditory stimulus / positive regulation of action potential / smooth muscle contraction involved in micturition / positive regulation of blood pressure / regulation of smooth muscle cell proliferation / positive regulation of hormone secretion / positive regulation of vascular permeability / positive regulation of ossification / regulation of smooth muscle cell migration / positive regulation of leukocyte migration / response to pain /  eating behavior / positive regulation of epithelial cell migration / behavioral response to pain / angiotensin-mediated drinking behavior / PKA activation in glucagon signalling / eating behavior / positive regulation of epithelial cell migration / behavioral response to pain / angiotensin-mediated drinking behavior / PKA activation in glucagon signalling /  associative learning / hair follicle placode formation / associative learning / hair follicle placode formation /  intracellular transport / intracellular transport /  D1 dopamine receptor binding / developmental growth / neuropeptide signaling pathway / D1 dopamine receptor binding / developmental growth / neuropeptide signaling pathway /  long-term memory / sperm flagellum / Hedgehog 'off' state / positive regulation of cAMP-mediated signaling / adenylate cyclase-activating adrenergic receptor signaling pathway / response to electrical stimulus / positive regulation of vasoconstriction / sensory perception of pain / positive regulation of stress fiber assembly / sperm midpiece / activation of adenylate cyclase activity / adenylate cyclase activator activity / cellular response to nerve growth factor stimulus / trans-Golgi network membrane / response to progesterone / response to nicotine / positive regulation of epithelial cell proliferation / positive regulation of synaptic transmission, GABAergic / G-protein beta/gamma-subunit complex binding / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / long-term memory / sperm flagellum / Hedgehog 'off' state / positive regulation of cAMP-mediated signaling / adenylate cyclase-activating adrenergic receptor signaling pathway / response to electrical stimulus / positive regulation of vasoconstriction / sensory perception of pain / positive regulation of stress fiber assembly / sperm midpiece / activation of adenylate cyclase activity / adenylate cyclase activator activity / cellular response to nerve growth factor stimulus / trans-Golgi network membrane / response to progesterone / response to nicotine / positive regulation of epithelial cell proliferation / positive regulation of synaptic transmission, GABAergic / G-protein beta/gamma-subunit complex binding / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor /  bone development / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / adenylate cyclase-activating G protein-coupled receptor signaling pathway / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK / Sensory perception of sweet, bitter, and umami (glutamate) taste / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / Adrenaline,noradrenaline inhibits insulin secretion / bone development / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / adenylate cyclase-activating G protein-coupled receptor signaling pathway / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK / Sensory perception of sweet, bitter, and umami (glutamate) taste / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / Adrenaline,noradrenaline inhibits insulin secretion /  platelet aggregation / Glucagon-type ligand receptors / platelet aggregation / Glucagon-type ligand receptors /  cognition / Vasopressin regulates renal water homeostasis via Aquaporins / positive regulation of GTPase activity / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / adenylate cyclase-activating dopamine receptor signaling pathway / ADP signalling through P2Y purinoceptor 1 / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / sensory perception of taste / GPER1 signaling / G-protein beta-subunit binding / cognition / Vasopressin regulates renal water homeostasis via Aquaporins / positive regulation of GTPase activity / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / adenylate cyclase-activating dopamine receptor signaling pathway / ADP signalling through P2Y purinoceptor 1 / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / sensory perception of taste / GPER1 signaling / G-protein beta-subunit binding /  heterotrimeric G-protein complex / Inactivation, recovery and regulation of the phototransduction cascade / heterotrimeric G-protein complex / Inactivation, recovery and regulation of the phototransduction cascade /  extracellular vesicle / G alpha (12/13) signalling events / signaling receptor complex adaptor activity / sensory perception of smell extracellular vesicle / G alpha (12/13) signalling events / signaling receptor complex adaptor activity / sensory perception of smellSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) / Homo sapiens (human) /   Lama glama (llama) Lama glama (llama) | |||||||||
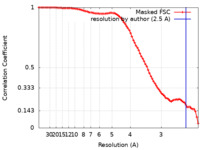
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.5 Å cryo EM / Resolution: 2.5 Å | |||||||||
 Authors Authors | Chen M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2024 Journal: Nat Methods / Year: 2024Title: Improving resolution and resolvability of single-particle cryoEM structures using Gaussian mixture models. Authors: Muyuan Chen / Michael F Schmid / Wah Chiu /  Abstract: Cryogenic electron microscopy is widely used in structural biology, but its resolution is often limited by the dynamics of the macromolecule. Here we developed a refinement protocol based on Gaussian ...Cryogenic electron microscopy is widely used in structural biology, but its resolution is often limited by the dynamics of the macromolecule. Here we developed a refinement protocol based on Gaussian mixture models that integrates particle orientation and conformation estimation and improves the alignment for flexible domains of protein structures. We demonstrated this protocol on multiple datasets, resulting in improved resolution and resolvability, locally and globally, by visual and quantitative measures. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41840.map.gz emd_41840.map.gz | 1.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41840-v30.xml emd-41840-v30.xml emd-41840.xml emd-41840.xml | 20.1 KB 20.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41840_fsc.xml emd_41840_fsc.xml | 8.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_41840.png emd_41840.png | 117.5 KB | ||
| Filedesc metadata |  emd-41840.cif.gz emd-41840.cif.gz | 6.3 KB | ||
| Others |  emd_41840_half_map_1.map.gz emd_41840_half_map_1.map.gz emd_41840_half_map_2.map.gz emd_41840_half_map_2.map.gz | 49 MB 49 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41840 http://ftp.pdbj.org/pub/emdb/structures/EMD-41840 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41840 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41840 | HTTPS FTP |
-Related structure data
| Related structure data |  8u26MC  8u28C  8u2cC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41840.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41840.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.146 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_41840_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
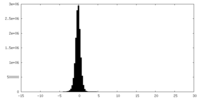
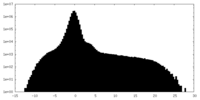
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_41840_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
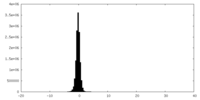
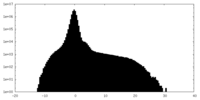
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Substance P bound to active human neurokinin 1 receptor in comple...
| Entire | Name: Substance P bound to active human neurokinin 1 receptor in complex with miniGs399 |
|---|---|
| Components |
|
-Supramolecule #1: Substance P bound to active human neurokinin 1 receptor in comple...
| Supramolecule | Name: Substance P bound to active human neurokinin 1 receptor in complex with miniGs399 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Reprocessing of EMPIAR-10786. Sample information is the same as EMD-24570, PDB-7RMH. |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 141 KDa |
-Macromolecule #1: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.907684 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NSKTEDQRNE EKAQREANKK IEKQLQKDKQ VYRATHRLLL LGADNSGKST IVKQMRILHG GSGGSGGTSG IFETKFQVDK VNFHMFDVG GQRDERRKWI QCFNDVTAII FVVDSSDYNR LQEALNLFKS IWNNRWLRTI SVILFLNKQD LLAEKVLAGK S KIEDYFPE ...String: NSKTEDQRNE EKAQREANKK IEKQLQKDKQ VYRATHRLLL LGADNSGKST IVKQMRILHG GSGGSGGTSG IFETKFQVDK VNFHMFDVG GQRDERRKWI QCFNDVTAII FVVDSSDYNR LQEALNLFKS IWNNRWLRTI SVILFLNKQD LLAEKVLAGK S KIEDYFPE FARYTTPEDA TPEPGEDPRV TRAKYFIRDE FLRISTASGD GRHYCYPHFT CAVDTENARR IFNDCRDIIQ RM HLRQYEL L UniProtKB: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.786566 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MHHHHHHLEV LFQGPEDQVD PRLIDGKGSS GSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIY AMHWGTDSRL LVSASQDGKL IIWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR E GNVRVSRE ...String: MHHHHHHLEV LFQGPEDQVD PRLIDGKGSS GSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIY AMHWGTDSRL LVSASQDGKL IIWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR E GNVRVSRE LAGHTGYLSC CRFLDDNQIV TSSGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KL WDVREGM CRQTFTGHES DINAICFFPN GNAFATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYD DFNCNV WDALKADRAG VLAGHDNRVS CLGVTDDGMA VATGSWDSFL KIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.56375 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFC UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Nanobody 35
| Macromolecule | Name: Nanobody 35 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Lama glama (llama) Lama glama (llama) |
| Molecular weight | Theoretical: 15.398067 KDa |
| Recombinant expression | Organism:   Lama glama (llama) Lama glama (llama) |
| Sequence | String: QVQLQESGGG LVQPGGSLRL SCAASGFTFS NYKMNWVRQA PGKGLEWVSD ISQSGASISY TGSVKGRFTI SRDNAKNTLY LQMNSLKPE DTAVYYCARC PAPFTRDCFD VTSTTYAYRG QGTQVTVSSG SEDQVDPRLI DGK |
-Macromolecule #5: Substance-P receptor
| Macromolecule | Name: Substance-P receptor / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.542867 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DYKDDDDASI DMDNVLPVDS DLSPNISTNT SEPNQFVQPA WQIVLWAAAY TVIVVTSVVG NVVVMWIILA HKRMRTVTNY FLVNLAFAE ASMAAFNTVV NFTYAVHNEW YYGLFYCKFH NFFPIAAVFA SIYSMTAVAF DRYMAIIHPL QPRLSATATK V VICVIWVL ...String: DYKDDDDASI DMDNVLPVDS DLSPNISTNT SEPNQFVQPA WQIVLWAAAY TVIVVTSVVG NVVVMWIILA HKRMRTVTNY FLVNLAFAE ASMAAFNTVV NFTYAVHNEW YYGLFYCKFH NFFPIAAVFA SIYSMTAVAF DRYMAIIHPL QPRLSATATK V VICVIWVL ALLLAFPQGY YSTTETMPSR VVCMIEWPEH PNKIYEKVYH ICVTVLIYFL PLLVIGYAYT VVGITLWASE IP GDSSDRY HEQVSAKRKV VKMMIVVVCT FAICWLPFHI FFLLPYINPD LYLKKFIQQV YLAIMWLAMS STMYNPIIYC CLN DRFRLG FKHAFRCCPF ISAGDYEGLE MKSTRYLQTQ GSVYKVSRLE TTISTVVGAH EEEPEDGPKA TPSSLDLTSN CSSR SDSKT MTESFSFSSN VLS UniProtKB: Substance-P receptor |
-Macromolecule #6: Protachykinin-1
| Macromolecule | Name: Protachykinin-1 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 1.350629 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: RPKPQQFFGL M UniProtKB: Protachykinin-1 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 30.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8u26: |
 Movie
Movie Controller
Controller


























 Z
Z Y
Y X
X