[English] 日本語
 Yorodumi
Yorodumi- EMDB-41629: Cryo-EM structure of the human MRS2 magnesium channel under Mg2+ ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human MRS2 magnesium channel under Mg2+ condition (C1 map) | |||||||||
 Map data Map data | local resolution filtered map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Magnesium / Magnesium /  Ion channel / Ion channel /  Membrane protein / METAL TRANSPORT / Ion Translocation / Divalent Ion / Membrane protein / METAL TRANSPORT / Ion Translocation / Divalent Ion /  Mg2+ / Mg2+ /  Pentamer Pentamer | |||||||||
| Function / homology | mitochondrial magnesium ion transmembrane transport / Magnesium transporter MRS2-like / Miscellaneous transport and binding events / magnesium ion transmembrane transporter activity / lactate metabolic process / transmembrane transport /  mitochondrial inner membrane / mitochondrial inner membrane /  mitochondrion / Magnesium transporter MRS2 homolog, mitochondrial mitochondrion / Magnesium transporter MRS2 homolog, mitochondrial Function and homology information Function and homology information | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
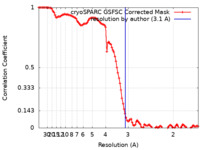
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.1 Å cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Lai LTF / Balaraman J / Zhou F / Matthies D | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structures of human magnesium channel MRS2 reveal gating and regulatory mechanisms Authors: Lai LTF / Balaraman J / Zhou F / Matthies D | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41629.map.gz emd_41629.map.gz | 4.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41629-v30.xml emd-41629-v30.xml emd-41629.xml emd-41629.xml | 26.9 KB 26.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41629_fsc.xml emd_41629_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_41629.png emd_41629.png | 77.1 KB | ||
| Masks |  emd_41629_msk_1.map emd_41629_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-41629.cif.gz emd-41629.cif.gz | 7 KB | ||
| Others |  emd_41629_additional_1.map.gz emd_41629_additional_1.map.gz emd_41629_additional_2.map.gz emd_41629_additional_2.map.gz emd_41629_half_map_1.map.gz emd_41629_half_map_1.map.gz emd_41629_half_map_2.map.gz emd_41629_half_map_2.map.gz | 62.3 MB 118 MB 116 MB 116 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41629 http://ftp.pdbj.org/pub/emdb/structures/EMD-41629 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41629 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41629 | HTTPS FTP |
-Related structure data
| Related structure data |  8tulC  8tupC C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41629.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41629.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | local resolution filtered map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_41629_msk_1.map emd_41629_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
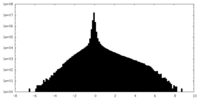
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: unsharpened map
| File | emd_41629_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
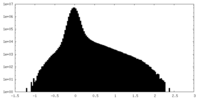
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: sharpened map
| File | emd_41629_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||
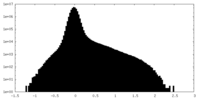
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_41629_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_41629_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of the pentameric human MRS2 magnesium channel ...
| Entire | Name: Cryo-EM structure of the pentameric human MRS2 magnesium channel under Mg2+ condition at an average resolution of 2.8 A, filtered to local resolution, C1 |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of the pentameric human MRS2 magnesium channel ...
| Supramolecule | Name: Cryo-EM structure of the pentameric human MRS2 magnesium channel under Mg2+ condition at an average resolution of 2.8 A, filtered to local resolution, C1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 219 KDa |
-Macromolecule #1: Mitochondrial RNA Splicing 2
| Macromolecule | Name: Mitochondrial RNA Splicing 2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MECLRSLPCL LPRAMRLPRR TLCALALDVT SVGPPVAACG RRANLIGRSR AAQLCGPDRL RVAGEVHRFR TSDVSQATLA SVAPVFTVT KFDKQGNVTS FERKKTELYQ ELGLQARDLR FQHVMSITVR NNRIIMRMEY LKAVITPECL LILDYRNLNL E QWLFRELP ...String: MECLRSLPCL LPRAMRLPRR TLCALALDVT SVGPPVAACG RRANLIGRSR AAQLCGPDRL RVAGEVHRFR TSDVSQATLA SVAPVFTVT KFDKQGNVTS FERKKTELYQ ELGLQARDLR FQHVMSITVR NNRIIMRMEY LKAVITPECL LILDYRNLNL E QWLFRELP SQLSGEGQLV TYPLPFEFRA IEALLQYWIN TLQGKLSILQ PLILETLDAL VDPKHSSVDR SKLHILLQNG KS LSELETD IKIFKESILE ILDEEELLEE LCVSKWSDPQ VFEKSSAGID HAEEMELLLE NYYRLADDLS NAARELRVLI DDS QSIIFI NLDSHRNVMM RLNLQLTMGT FSLSLFGLMG VAFGMNLESS LEEDHRIFWL ITGIMFMGSG LIWRRLLSFL GRQL EAPLP PMMASLPKKT LLADRSMELK NSLRLDGLGS GRSILTNRDY KDDDDK UniProtKB: Magnesium transporter MRS2 homolog, mitochondrial |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.3 Component:
Details: 20 mM HEPES, 150 mM NaCl, 40 mM MgCl2, 0.003% LMNG, pH 7.3 | ||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | ||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: LEICA EM GP Details: 400-mesh R1.2/1.3 Cu grids (Quantifoil) were made hydrophilic by glow discharging for 60 seconds with a current of 15 mA in a PELCO easiGlow system. The cryo grids were produced using a ...Details: 400-mesh R1.2/1.3 Cu grids (Quantifoil) were made hydrophilic by glow discharging for 60 seconds with a current of 15 mA in a PELCO easiGlow system. The cryo grids were produced using a Leica EM GP2 (Leica). The chamber was kept at 4 C and set to 95% humidity. 3 microliter sample at 0.5 mg/ml was applied to a glow-discharged holey grid, blotted for 6 s, and plunge frozen into liquid ethane and stored in liquid nitrogen.. | ||||||||||
| Details | This sample was monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 105000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 105000 |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV Details: Cryo-EM datasets were acquired with SerialEM using a Titan Krios (FEI, now ThermoFisher Scientific) operated at 300 keV and equipped with an energy filter and K3 camera (Gatan Inc.). Movies ...Details: Cryo-EM datasets were acquired with SerialEM using a Titan Krios (FEI, now ThermoFisher Scientific) operated at 300 keV and equipped with an energy filter and K3 camera (Gatan Inc.). Movies of 50 frames with a dose of 1 e-/A2 per frame (50 e-/A2 total dose) were recorded at a nominal magnification of 105,000x, corresponding to a physical pixel size of 0.83 A/px (super-resolution pixel size 0.415 A/px) in CDS mode at a dose rate of 10 e-/px/s and a defocus range of -0.7 to -2.0 um. In total, 3,991 and 9,656 movies were collected. |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 3991 / Average exposure time: 3.462 sec. / Average electron dose: 50.0 e/Å2 Details: Cryo-EM datasets were acquired with SerialEM using a Titan Krios (FEI, now ThermoFisher Scientific) operated at 300 keV and equipped with an energy filter and K3 camera (Gatan Inc.). Movies ...Details: Cryo-EM datasets were acquired with SerialEM using a Titan Krios (FEI, now ThermoFisher Scientific) operated at 300 keV and equipped with an energy filter and K3 camera (Gatan Inc.). Movies of 50 frames with a dose of 1 e-/A2 per frame (50 e-/A2 total dose) were recorded at a nominal magnification of 105,000x, corresponding to a physical pixel size of 0.83 A/px (super-resolution pixel size 0.415 A/px) in CDS mode at a dose rate of 10 e-/px/s and a defocus range of -0.7 to -2.0 um. In total, 3,991 and 9,656 movies were collected. |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Details | The model was then manually rebuilt in COOT v.0.9.7 using the local resolution filtered map, which was generated from refinement with C5 applied. The Mg2+ ions assigned in the pore regions were confirmed in the C1 map. Loop regions (residues 174-181, 273-287) were built with the unsharpened map. Iterative rounds of manual refinement in COOT and real-space refinement in Phenix v.1.20.1-4487 were performed. |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 141 |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X