[English] 日本語
 Yorodumi
Yorodumi- EMDB-40827: Bovine multidrug resistance protein 4 (MRP4) bound to prostagland... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bovine multidrug resistance protein 4 (MRP4) bound to prostaglandin E1 in MSP lipid nanodisc | |||||||||
 Map data Map data | Output density from cryoSPARC non-uniform refinement | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ABC transporter / multidrug resistance-associated protein / membrane protein / transport protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationParacetamol ADME / Platelet degranulation / Azathioprine ADME / ABC-family proteins mediated transport / ATPase-coupled transmembrane transporter activity / ABC-type transporter activity / transmembrane transport / ATP hydrolysis activity / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Pourmal S / Stroud RM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Structural basis of prostaglandin efflux by MRP4. Authors: Sergei Pourmal / Evan Green / Ruchika Bajaj / Ilan E Chemmama / Giselle M Knudsen / Meghna Gupta / Andrej Sali / Yifan Cheng / Charles S Craik / Deanna L Kroetz / Robert M Stroud /  Abstract: Multidrug resistance protein 4 (MRP4) is a broadly expressed ATP-binding cassette transporter that is unique among the MRP subfamily for transporting prostanoids, a group of signaling molecules ...Multidrug resistance protein 4 (MRP4) is a broadly expressed ATP-binding cassette transporter that is unique among the MRP subfamily for transporting prostanoids, a group of signaling molecules derived from unsaturated fatty acids. To better understand the basis of the substrate selectivity of MRP4, we used cryogenic-electron microscopy to determine six structures of nanodisc-reconstituted MRP4 at various stages throughout its transport cycle. Substrate-bound structures of MRP4 in complex with PGE, PGE and the sulfonated-sterol DHEA-S reveal a common binding site that accommodates a diverse set of organic anions and suggest an allosteric mechanism for substrate-induced enhancement of MRP4 ATPase activity. Our structure of a catalytically compromised MRP4 mutant bound to ATP-Mg is outward-occluded, a conformation previously unobserved in the MRP subfamily and consistent with an alternating-access transport mechanism. Our study provides insights into the endogenous function of this versatile efflux transporter and establishes a basis for MRP4-targeted drug design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40827.map.gz emd_40827.map.gz | 97.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40827-v30.xml emd-40827-v30.xml emd-40827.xml emd-40827.xml | 21.1 KB 21.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40827.png emd_40827.png | 67.9 KB | ||
| Filedesc metadata |  emd-40827.cif.gz emd-40827.cif.gz | 7.3 KB | ||
| Others |  emd_40827_additional_1.map.gz emd_40827_additional_1.map.gz emd_40827_half_map_1.map.gz emd_40827_half_map_1.map.gz emd_40827_half_map_2.map.gz emd_40827_half_map_2.map.gz | 91.2 MB 95.6 MB 95.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40827 http://ftp.pdbj.org/pub/emdb/structures/EMD-40827 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40827 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40827 | HTTPS FTP |
-Validation report
| Summary document |  emd_40827_validation.pdf.gz emd_40827_validation.pdf.gz | 823.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40827_full_validation.pdf.gz emd_40827_full_validation.pdf.gz | 822.8 KB | Display | |
| Data in XML |  emd_40827_validation.xml.gz emd_40827_validation.xml.gz | 13.1 KB | Display | |
| Data in CIF |  emd_40827_validation.cif.gz emd_40827_validation.cif.gz | 15.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40827 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40827 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40827 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40827 | HTTPS FTP |
-Related structure data
| Related structure data |  8sx8MC  8swnC  8sx7C  8sx9C  8sxaC  8sxbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40827.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40827.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Output density from cryoSPARC non-uniform refinement | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.835 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Output density from DeepEMhancer. Inputs are two half...
| File | emd_40827_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Output density from DeepEMhancer. Inputs are two half maps from cryoSPARC non-uniform refinement, using the "high resolution" protocol. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A from cryoSPARC non-uniform refinement.
| File | emd_40827_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A from cryoSPARC non-uniform refinement. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B from cryoSPARC non-uniform refinement.
| File | emd_40827_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B from cryoSPARC non-uniform refinement. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Bovine multidrug resistance protein 4 (MRP4) E1202Q mutant bound ...
| Entire | Name: Bovine multidrug resistance protein 4 (MRP4) E1202Q mutant bound to ATP in MSP lipid nanodisc |
|---|---|
| Components |
|
-Supramolecule #1: Bovine multidrug resistance protein 4 (MRP4) E1202Q mutant bound ...
| Supramolecule | Name: Bovine multidrug resistance protein 4 (MRP4) E1202Q mutant bound to ATP in MSP lipid nanodisc type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 150 KDa |
-Macromolecule #1: Multidrug resistance-associated protein 4
| Macromolecule | Name: Multidrug resistance-associated protein 4 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 149.587297 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQPVYPEVKP NPLRNANLCS RIFFWWLNPL FKIGHKRRLE EDDMYSVLPE DRSQHLGEEL QGYWDQEVLR AEKDAREPSL TKAIIKCYW KSYVVLGIFT LIEESTRVVQ PIILGKIIGY FENYDPSDSA ALYEAHGYAG VLSACTLVLA ILHHLYFYHV Q CAGMRLRV ...String: MQPVYPEVKP NPLRNANLCS RIFFWWLNPL FKIGHKRRLE EDDMYSVLPE DRSQHLGEEL QGYWDQEVLR AEKDAREPSL TKAIIKCYW KSYVVLGIFT LIEESTRVVQ PIILGKIIGY FENYDPSDSA ALYEAHGYAG VLSACTLVLA ILHHLYFYHV Q CAGMRLRV AMCHMIYRKA LRLSNSAMGK TTTGQIVNLL SNDVNKFDQV TIFLHFLWAG PLQAIVVTAL LWMEIGISCL AG MAVLIIL LPLQSCIGKL FSSLRSKTAA FTDTRIRTMN EVITGIRIIK MYAWEKSFAD LITNLRRKEI SKILRSSYLR GMN LASFFV ASKIIVFVTF TTYVFLGNVI TASRVFVAVS LYGAVRLTVT LFFPSAVEKV SEAFVSIRRI KNFLLLDEIT QLHS QLPSD GKMIVNVQDF TAFWDKASDT PTLQSLSFTV RPGELLAVVG PVGAGKSSLL SAVLGELPPN QGQVSVHGRI AYVSQ QPWV FSGTVRSNIL FGKKYEKERY EKVIKACALK KDLQLLEDGD LTMIGDRGTT LSGGQKARVN LARAVYQDAD IYLLDD PLS AVDAEVSRHL FELCICQALH EKIRILVTHQ LQYLKAASQI LILKDGQMVQ KGTYTEFLKS GIDFGSLLKK ENEEAEP SP VPGSPTLRNR TFSESSVWSQ QSSRPSLKEA TPEGQDTENI QVTLTEESRS EGKVGFKAYK NYFTAGAHWF IIIFLILV N LAAQVSYILQ DWWLSYWANQ QSALNVTVNG QGNVTEKLDL NWYLGIYSGL TASTVLFGIV RSLLVFFVLV SSSQTLHNQ MFESILRAPV LFFDRNPIGR ILNRFSKDIG HMDDLLPLTY LDFIQTFLQV IGVVGVAVAV IPWIAIPLVP LGIVFFVLRR YFLETSRDV KRLESTTRSP VFSHLSSSLQ GLWTIRAYKA EQRFQELFDS HQDLHSEAWF LFLTTSRWFA VRLDAICAVF V IVVAFGSL ILAKTLDAGQ VGLALSYALT LMGMFQWCVR QSAEVENMMI SVERVIEYTD LEKEAPWEYQ KRPLPSWPHE GV IIFDNVN FSYSLDGPLV LKHLTALIKS KEKVGIVGRT GAGKSSLIAA LFRLSEPEGK IWIDKILTTE IGLHDLRKKM SII PQEPVL FTGTMRKNLD PFNEHSDEEL WNALEEVQLK EAIEDLPGKM DTELAESGSN FSVGQRQLVC LARAILRKNR ILII DEATA NVDPRTDELI QKKIREKFAH CTVLTIAHRL NTIIDSDKIM VLDSGRLKEY DEPYVLLQNR DSLFYKMVQQ LGKAE AAAL TETAKQVYFK RNYPDITHNG HVVMNASSGQ PSAFTIFETA L UniProtKB: ATP binding cassette subfamily C member 4 |
-Macromolecule #2: 7-[(1R,3R)-3-hydroxy-2-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-5-oxocycl...
| Macromolecule | Name: 7-[(1R,3R)-3-hydroxy-2-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-5-oxocyclopentyl]heptanoic acid type: ligand / ID: 2 / Number of copies: 1 / Formula: XPG |
|---|---|
| Molecular weight | Theoretical: 354.481 Da |
| Chemical component information | 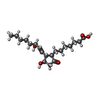 ChemComp-XPG: |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 1 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM Tris-HCL, 150 mM NaCl, 1 mM TCEP, 200 uM prostaglandin E1, 2% DMSO |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
| Details | Multidrug resistance-associated protein (MRP4) in MSP lipid nanodisc |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 8253 / Average electron dose: 66.0 e/Å2 Details: 7219 images collected with 3x record per hole. Merged with 1034 images collected 1x record per hole with 35 degree tilt. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X

























