[English] 日本語
 Yorodumi
Yorodumi- EMDB-40268: CryoEM map of a de novo designed T=4 octahedral nanocage hierarch... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM map of a de novo designed T=4 octahedral nanocage hierarchically built from pseudosymmetric trimers; design Oct(T=4)-3 | |||||||||
 Map data Map data | sharpened map using DeepEMhancer | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | icosahedral nanocage / T=4 / T=4 icosahedra /  nanomaterial / computational design / nanomaterial / computational design /  de novo / de novo /  DE NOVO PROTEIN DE NOVO PROTEIN | |||||||||
| Biological species | synthetic construct (others) | |||||||||
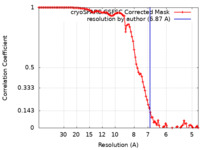
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 6.87 Å cryo EM / Resolution: 6.87 Å | |||||||||
 Authors Authors | Philomin A / Borst AJ / Kibler RD | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Design of four component T=4 tetrahedral, octahedral, and icosahedral protein nanocages through programmed symmetry breaking Authors: Lee S / Kibler RD / Hsia Y / Borst AJ / Philomin A / Kennedy MA / Stoddard B / Baker D | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40268.map.gz emd_40268.map.gz | 92 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40268-v30.xml emd-40268-v30.xml emd-40268.xml emd-40268.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40268_fsc.xml emd_40268_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_40268.png emd_40268.png | 134.6 KB | ||
| Filedesc metadata |  emd-40268.cif.gz emd-40268.cif.gz | 4.8 KB | ||
| Others |  emd_40268_additional_1.map.gz emd_40268_additional_1.map.gz emd_40268_half_map_1.map.gz emd_40268_half_map_1.map.gz emd_40268_half_map_2.map.gz emd_40268_half_map_2.map.gz | 50.9 MB 95.4 MB 95.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40268 http://ftp.pdbj.org/pub/emdb/structures/EMD-40268 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40268 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40268 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40268.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40268.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map using DeepEMhancer | ||||||||||||||||||||
| Voxel size | X=Y=Z: 2.36 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: unsharpened map from final Non-Uniform Refinement
| File | emd_40268_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map from final Non-Uniform Refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A from final Non-Uniform Refinement
| File | emd_40268_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A from final Non-Uniform Refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B from final Non-Uniform Refinement
| File | emd_40268_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B from final Non-Uniform Refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Oct(T=4)-3
| Entire | Name: Oct(T=4)-3 |
|---|---|
| Components |
|
-Supramolecule #1: Oct(T=4)-3
| Supramolecule | Name: Oct(T=4)-3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: A hierarchically designed T=4 octahedral nanocage |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: Oct(T=4)-3 chain A
| Macromolecule | Name: Oct(T=4)-3 chain A / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MGSLELALKA LQILVNAAYV LAEIARDRGN EELLEKAARL AEEAARQAER IARQARKEGN LELALKALQI LVNAAYVLAE IARDRGNEEL LEYAARLAEE AARQAIEIWA QAMEEGNQQL RTKAAHIILR AAEVLLEIAR DRGNQELLEK AASLVDAVAA LQQAAAAILE ...String: MGSLELALKA LQILVNAAYV LAEIARDRGN EELLEKAARL AEEAARQAER IARQARKEGN LELALKALQI LVNAAYVLAE IARDRGNEEL LEYAARLAEE AARQAIEIWA QAMEEGNQQL RTKAAHIILR AAEVLLEIAR DRGNQELLEK AASLVDAVAA LQQAAAAILE GDVEKAVRAA QEAVKAAKEA GDNDMLRAVA IAALRIAKEA EKQGNVEVAV KAARVAVEAA KQAGDQDVLL KVATQAARIA LEAIKQGNLE VKLEALKVAQ EALKQTGGSG GSHHHHHH |
-Macromolecule #2: Oct(T=4)-3 chain B
| Macromolecule | Name: Oct(T=4)-3 chain B / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Sequence | String: MGSPRLVLRA LENMVRAAHT LAEIARDNGN EEWLERAARL AEEVARRAER LAREARKEGN LELALKALQI LVNAAYVLAE IARDRGNEEE LEYAARLAEE AARQAIEIAA QAMEEGNLEL ALKALQIIVN AAYVLAEIAR DRGNEELLEK AASLAEAAAA LAEAIAAILE ...String: MGSPRLVLRA LENMVRAAHT LAEIARDNGN EEWLERAARL AEEVARRAER LAREARKEGN LELALKALQI LVNAAYVLAE IARDRGNEEE LEYAARLAEE AARQAIEIAA QAMEEGNLEL ALKALQIIVN AAYVLAEIAR DRGNEELLEK AASLAEAAAA LAEAIAAILE GDVEKAVRAA QEAVKAAKEA GDNDMLRAVA IAALRIAKEA EKQGNVEVAV KAARVAVEAA KQAGDQDVLK KVAAQAARIM LEAIKQGNTE VALEALKVAQ EALKQTGGSG GSHHHHHH |
-Macromolecule #3: Oct(T=4)-3 chain C
| Macromolecule | Name: Oct(T=4)-3 chain C / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MGSPELFLQD LRSLVEAARI LARLARQRGD EHALERAARW AEQAARQAER LARQARKEGN LELALKALQI LVNAAYVLAE IARDRGNEEL LEYAARLAEE AARQAIEIAA QAMEEGNFEL ALEALEIINE AARVLARIAH HRGNQELLEK AASLTHASAA LSRAIAAILE ...String: MGSPELFLQD LRSLVEAARI LARLARQRGD EHALERAARW AEQAARQAER LARQARKEGN LELALKALQI LVNAAYVLAE IARDRGNEEL LEYAARLAEE AARQAIEIAA QAMEEGNFEL ALEALEIINE AARVLARIAH HRGNQELLEK AASLTHASAA LSRAIAAILE GDVEKAVRAA QEAVKAAKEA GDNDMLRAVA IAALRIAKEA EKQGNVEVAV KAARVAVEAA KQAGDNDVLR KVAEQALRIA KEAEKQGNVI VAMKAIDVAV EAAGQAGDID VIKKVADQTQ RIVDEWLKQG |
-Macromolecule #4: Oct(T=4)-3 chain ho
| Macromolecule | Name: Oct(T=4)-3 chain ho / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MGSPELFLQD LRSLVEAARI LARLARQRGD EHALERAARW AEQAARQAER LARQARKEGN LELALKALQI LVNAAYVLAE IARDRGNEEL LEYAARLAEE AARQAIEIWA QAMEEGNQQL RTKAAHIILR AAEVLLEIAR DRGNQELLEK AASLVDAVAA LQQAAAAILE ...String: MGSPELFLQD LRSLVEAARI LARLARQRGD EHALERAARW AEQAARQAER LARQARKEGN LELALKALQI LVNAAYVLAE IARDRGNEEL LEYAARLAEE AARQAIEIWA QAMEEGNQQL RTKAAHIILR AAEVLLEIAR DRGNQELLEK AASLVDAVAA LQQAAAAILE GDVEKAVRAA QEAVKAAKEA GDNDMLRAVA IAALRIAKEA EKQGNVEVAV KAARVAVEAA KQAGDNDVLR KVAEQALRIA KEAEKQGNVY VAAKAVQVAA EAAKQAGDID VLKKVIGQTQ RIRDEWVKQG GSGGSHHHHH H |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Details: 25mM Tris, 300mM NaCl |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.7000000000000001 µm Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.7000000000000001 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X