Entry Database : EMDB / ID : EMD-37663Title CryoEM structure of ZIKV rsNS1 Virus : Protein or peptide : Non-structural protein 1Ligand : 2-acetamido-2-deoxy-beta-D-glucopyranose / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Method / / Resolution : 3.0 Å Chew BLA / Luo D Funding support Organization Grant number Country Ministry of Education (MoE, Singapore) T2EP30220-0020
Journal : Npj Viruses / Year : 2024Title : Structural basis of Zika virus NS1 multimerization and human antibody recognitionAuthors : Chew BLA / Ngoh AQ / Phoo WW / Weng MJG / Sheng HJ / Chan KWK / Tan EYJ / Gelbart T / Xu C / Tan GS / Vasudevan SG / Luo D History Deposition Oct 5, 2023 - Header (metadata) release May 8, 2024 - Map release May 8, 2024 - Update May 8, 2024 - Current status May 8, 2024 Processing site : PDBj / Status : Released
Show all Show less
 Open data
Open data Basic information
Basic information
 Map data
Map data Sample
Sample Keywords
Keywords Flavivirus /
Flavivirus /  ZIKA / NS1 /
ZIKA / NS1 /  VIRAL PROTEIN
VIRAL PROTEIN Function and homology information
Function and homology information flavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity /
flavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity /  viral capsid / nucleoside-triphosphate phosphatase /
viral capsid / nucleoside-triphosphate phosphatase /  double-stranded RNA binding / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of TBK1 activity / symbiont-mediated suppression of host toll-like receptor signaling pathway / mRNA (guanine-N7)-methyltransferase ...symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity /
double-stranded RNA binding / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of TBK1 activity / symbiont-mediated suppression of host toll-like receptor signaling pathway / mRNA (guanine-N7)-methyltransferase ...symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity /  flavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity /
flavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity /  viral capsid / nucleoside-triphosphate phosphatase /
viral capsid / nucleoside-triphosphate phosphatase /  double-stranded RNA binding / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of TBK1 activity / symbiont-mediated suppression of host toll-like receptor signaling pathway / mRNA (guanine-N7)-methyltransferase / methyltransferase cap1 / clathrin-dependent endocytosis of virus by host cell /
double-stranded RNA binding / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of TBK1 activity / symbiont-mediated suppression of host toll-like receptor signaling pathway / mRNA (guanine-N7)-methyltransferase / methyltransferase cap1 / clathrin-dependent endocytosis of virus by host cell /  mRNA (nucleoside-2'-O-)-methyltransferase activity / mRNA 5'-cap (guanine-N7-)-methyltransferase activity /
mRNA (nucleoside-2'-O-)-methyltransferase activity / mRNA 5'-cap (guanine-N7-)-methyltransferase activity /  RNA helicase activity / host cell endoplasmic reticulum membrane / host cell perinuclear region of cytoplasm /
RNA helicase activity / host cell endoplasmic reticulum membrane / host cell perinuclear region of cytoplasm /  protein dimerization activity /
protein dimerization activity /  RNA helicase / induction by virus of host autophagy /
RNA helicase / induction by virus of host autophagy /  RNA-directed RNA polymerase / viral RNA genome replication /
RNA-directed RNA polymerase / viral RNA genome replication /  RNA-dependent RNA polymerase activity / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane /
RNA-dependent RNA polymerase activity / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane /  viral envelope /
viral envelope /  lipid binding / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / host cell nucleus / virion attachment to host cell / GTP binding / virion membrane / structural molecule activity /
lipid binding / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / host cell nucleus / virion attachment to host cell / GTP binding / virion membrane / structural molecule activity /  ATP hydrolysis activity /
ATP hydrolysis activity /  proteolysis / extracellular region /
proteolysis / extracellular region /  ATP binding /
ATP binding /  membrane /
membrane /  metal ion binding
metal ion binding

 Zika virus
Zika virus single particle reconstruction /
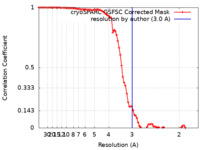
single particle reconstruction /  cryo EM / Resolution: 3.0 Å
cryo EM / Resolution: 3.0 Å  Authors
Authors Singapore, 1 items
Singapore, 1 items  Citation
Citation Journal: Npj Viruses / Year: 2024
Journal: Npj Viruses / Year: 2024 Structure visualization
Structure visualization Downloads & links
Downloads & links emd_37663.map.gz
emd_37663.map.gz EMDB map data format
EMDB map data format emd-37663-v30.xml
emd-37663-v30.xml emd-37663.xml
emd-37663.xml EMDB header
EMDB header emd_37663_fsc.xml
emd_37663_fsc.xml FSC data file
FSC data file emd_37663.png
emd_37663.png emd-37663.cif.gz
emd-37663.cif.gz emd_37663_half_map_1.map.gz
emd_37663_half_map_1.map.gz emd_37663_half_map_2.map.gz
emd_37663_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-37663
http://ftp.pdbj.org/pub/emdb/structures/EMD-37663 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37663
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37663




 F&H Search
F&H Search Links
Links EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource Map
Map Download / File: emd_37663.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
Download / File: emd_37663.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Sample components
Sample components

 Zika virus
Zika virus
 Homo sapiens (human)
Homo sapiens (human)

 Zika virus
Zika virus
 Homo sapiens (human)
Homo sapiens (human)
 cryo EM
cryo EM Processing
Processing single particle reconstruction
single particle reconstruction Sample preparation
Sample preparation Electron microscopy
Electron microscopy FIELD EMISSION GUN
FIELD EMISSION GUN
 Image processing
Image processing
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)